Research article
PCR Optimization Prior to Genetic
Diversity Assessment of Sesame (Sesamum indicum L.) Genotypes Using
Inter-Primer Binding Site (IPBS) Markers
Seval
ELİŞ *,1,3,
Büşra Polat2,
Ferhat KIZILGEÇİ1, Aras
TÜRKOĞLU4, Mehmet Yıldırım3
1Mardin Artuklu
University, Kızıltepe Faculty of Agricultural Sciences and Technologies,
Department of Field Crops, Mardin, Türkiye
2Department of Field
Crops, Faculty of Agriculture, Ataturk University, Erzurum 25240, Türkiye
3Dicle University, Faculty of Agriculture, Department of
Field Crops, 21280, Sur, Diyarbakır, Türkiye
4Necmettin
Erbakan University, Faculty of Agriculture, Department of Field Crops, , Konya
42310, Türkiye
Received: 23-11-2025 Accepted: 06-12-2025 Published online: 29-12-2025
DOI: https://doi.org/10.33687/ricosbiol.03.012.94
Abstract
This
study aimed to accurately and reliably determine the genetic diversity among
sesame (Sesamum indicum L.) genotypes, which is an important oil crop.
To achieve this, Polymerase Chain Reaction (PCR) conditions based on
Inter-Primer Binding Site (IPBS) molecular markers were optimized to reveal
genetic variation, which forms the basis of plant breeding programs. For the
methodology, DNA was isolated from fresh sesame leaves grown under controlled
conditions using the CTAB method and analyzed from 50 local sesame lines. Since
PCR success is directly dependent on the specificity of the primers and
reaction parameters like temperature, the gradient temperature PCR method was
applied using 22 different iPBS primers to determine the optimal annealing
temperatures. According to the findings, 20 out of the 22 primers successfully
generated polymorphic bands, revealing genetic diversity. Determining the
optimal PCR conditions was critical for identifying the binding temperature at
which iPBS primers exhibited the highest polymorphism. For example, Primer 2277
showed high amplification and activity at 47.6 and 50.9°C, while Primer 2218
was highly active at 50.9°C. This optimization establishes a precise molecular
foundation that will contribute to future sesame breeding programs.
Editor’s
Note: This manuscript was selected for
publication in Ricos Biology Journal from the World Conference
on Sustainable Life Sciences (WOCOLS 2025) held in Mardin, Türkiye. Ricos
Biology Journal served as a media partner for the conference.
![]()
Keywords:
local
sesame lines, IPBS, optimal annealing temperatures, molecular marker
.
I. Introduction
Sesame (Sesamum indicum L.) is an important oil crop
worldwide, with seeds containing high amounts of oil and protein (Saxena and Bisen,
2017). Its tolerance to drought and
other abiotic stresses increases sesame's potential for cultivation in marginal lands (Lakhanpaul et al., 2012). The
use of sesame plants, particularly local lines, in breeding programs and the genetic
diversity they offer is important. Identifying genetic diversity, which forms the
basis of plant breeding programs, enables the development
of new varieties with superior traits.
Molecular markers, one of the powerful
tools used for this purpose,
play an important role in assessing genetic diversity.
Inter-Primer Binding Site
(IPBS) markers, which are molecular markers,
are linked to retrotransposon regions to reveal
genetic variation (Özer et
al., 2017). High levels of polymorphism
(genetic diversity) can be detected because IPBS markers target recurrent DNA sequences in the plant genome.
Polymerase chain reaction (PCR) is a fundamental molecular biology method for amplifying
specific DNA regions (Al-Hadeithi and Jasim,
2021). The specificity of the primers used
and the reaction
parameters, particularly temperature, directly affect PCR success. Therefore, PCR optimization is
a critical step to ensure the accurate
and efficient amplification of target DNA regions.
Molecular markers are
one of the powerful tools used for this
purpose and play an important role in assessing genetic diversity. Inter-Primer Binding
Site (IPBS) markers, which are molecular markers
associated with retrotransposon regions and reveal genetic
variation, enable the detection of high levels of polymorphism (genetic diversity) because they target repetitive DNA sequences in the plant genome.
The aim of this
study was to determine the
ideal annealing temperatures
for various iPBS primers and
to optimize PCR analysis based on iPBS markers
in sesame genotypes. The resulting optimization
is expected to form a basis for a precise
and reliable assessment of the genetic diversity of sesame genotypes. The resulting optimization
is expected to form a basis for the
precise and reliable assessment of the genetic diversity
of sesame genotypes.
Material and Methods
In this study,
DNA samples from 50 different local sesame lines were
used for temperature detection. For this study,
fresh leaves of sesame were used
in a plant growth chamber under controlled
conditions.
DNA Isolation
Freshly harvested 3-5 leaves (100-150 mg)
from plants grown in a growth chamber were ground
into powder using liquid nitrogen.
DNA extraction was performed according to the CTAB method
developed by Doyle and Doyle (1990).
The sample was mixed
with CTAB buffer (2% CTAB)
20 g CTAB/L, 20 mM EDTA, 100 mM
Tris-HCl pH 8.0, 1.4 M NaCl, 0.2% Mercaptoethanol)
and was vortexed
after adding one unit of proteinase
K to each sample. The samples
were incubated for 60 min at 60°C; after the incubation,
1 U (20 mg/ml) proteinase K was
added to the mixture. Added
equal volume of chloroform/isoamylalcohol (24:1),
mixed for about 15 min, then
the samples were subjected to centrifugation at 16,000 xg for 10 min,
and supernatants were transferred to new centrifuge
tubes. Precipitated DNA with 2/3 volume of cold isopropanol centrifuged at 10,000 xg for 20 min. Supernatants were discarded, pellets were dried
at room temperature and added to
1X TE buffer, and then left overnight
at 4°C. The samples were incubated for 3 h at 65°C and 20 μL of RNase A (10 mg/mL) was added
and then incubated for 1 h again. The upper
phase was transferred to a new tube that
contained 100 l of 1.2 M NaCl. After
adding 0.6 ml of isopropanol,
it was left to stand for
1 hour at -20°C. Samples taken from -20°C were centrifuged at 6000 rpm for 10 minutes at +4°C, and the liquid
portion was removed by allowing
the DNA to settle at the bottom
of the tubes. The pellet was
dried and then carefully washed with 500 ml of 70% ethanol
without damaging the pellet. The
DNA was then dissolved in 100 µl of water.
For the qualitative analysis of DNA samples, a 0.8% agarose gel was prepared using 1X SB buffer. To load
the samples into the gel wells,
5 µl of DNA sample, 5 µl of ddH₂O,
and 3 µl of 6X loading buffer were mixed
to prepare a total mixture of 13 µl. This mixture was carefully
placed into the wells formed
in the gel. Gel electrophoresis
was performed at 70 V for 90 minutes, and after the
process was completed, the DNA bands were visualized
using a UV device (Vilber Lourmat, France).
The concentrations of the obtained DNA samples were measured using
a Quantus Fluorometer (Invitrogen, USA) by taking 2 µl from each sample. The
measurements were performed according to the QuantiFluor™
ONE dsDNA System Kit protocol.
PCR
Components and Conditions
Twenty-two iPBS markers were
used to determine
primer temperatures. These primers were selected
because they demonstrated high polymorphism and efficiency in various plant species
in previous studies (Özer et
al., 2017; Erkoç et al., 2024). Only 20 of
the 22 primers produced amplification products at the appropriate temperature. The PCR procedure was performed using
a MultiGene Gradient Thermal Cycler (TC9600-G-230V, Labnet International, Inc.). For the gradient
PCR test, reagents were added to 0.2 ml PCR tubes at the ratios
shown in Table (1).
Table (1): Mixing
ratios and final reaction volume of PCR components.
|
Component |
Amount |
|
10X PCR Buffer |
2 μL |
|
10 mM Dntp |
2 μL |
|
25 mM MgCl2 |
0.5 μL |
|
Taq DNA polymerase |
0.5 μL |
|
100 pmol primer |
1 μL |
Table (2): Names and 5'→3' sequences of the 22 different iPBS primers used in gradient PCR analyses.
|
Primer No |
IPBS Primer |
Sequence 5-3’ |
|
1 |
2077 |
CTCACGATGCCA |
|
2 |
2218 |
CTCCAGCTCCGATTACCA |
|
3 |
2226 |
CGGTGACCTTTGATACCA |
|
4 |
2228 |
CATTGGCTCTTGATACCA |
|
5 |
2230 |
TCTAGGCGTCTGATACCA |
|
6 |
2232 |
AGAGAGGCTCGGATACCA |
|
7 |
2237 |
CCCCTACCTGGCGTGCCA |
|
8 |
2243 |
AGTCAGGCTCTGTTACCA |
|
9 |
2244 |
GGAAGGCTCTGATTACCA |
|
10 |
2246 |
ACTAGGCTCTGTATACCA |
|
11 |
2249 |
AACCGACCTCTGATACCA |
|
12 |
2251 |
GAACAGGCGATGATACCA |
|
13 |
2252 |
TCATGGCTCATGATACCA |
|
14 |
2253 |
TCGAGGCTCTAGATACCA |
|
15 |
2272 |
GGCTCAGATGCCA |
|
16 |
2277 |
GGCGATGATACCA |
|
17 |
2379 |
TCCAGAGATCCA |
|
18 |
2380 |
CAACCTGATCCA |
|
19 |
2389 |
ACATCCTTCCCA |
|
20 |
2393 |
TACGGTACGCCA |
|
21 |
2401 |
AGTTAAGCTTTGATACCA |
|
22 |
2402 |
TCTAAGCTCTTGATACCA |
Results
In this study, PCR optimisation based on iPBS markers was
performed to determine the genetic
diversity of local sesame lines. Twenty
of the 22 primers used in the study
produced polymorphic bands. The primers
generally showed effective amplification within a specific temperature range. Amplification efficiency decreased at temperatures outside this range.
Some primers produced strong and distinct bands,
while others produced weak or
no amplification products. Some primers showed high polymorphism at a specific temperature in both DNA samples. Primer 2277 at
47.6°C and 50.9°C; Primer 2218 at 50.9°C; Primer 2246
at 45°C; Primer 2393 at 52.8°C; and Primer 2251 at
56.5°C showed the highest amplification and activity. Primers
2266, 2244, 2252, 2243, and 2401 showed
high activity in the 47-48.3°C range; Primers 2402 and 2077 at 49.4°C; and Primers 2237 and 2249 in the 51.1-51.4°C range. Primers 2379 and 2389, however, yielded weak or
no amplification products. Gel images of the amplification products of the primers are presented
in Figure (1).
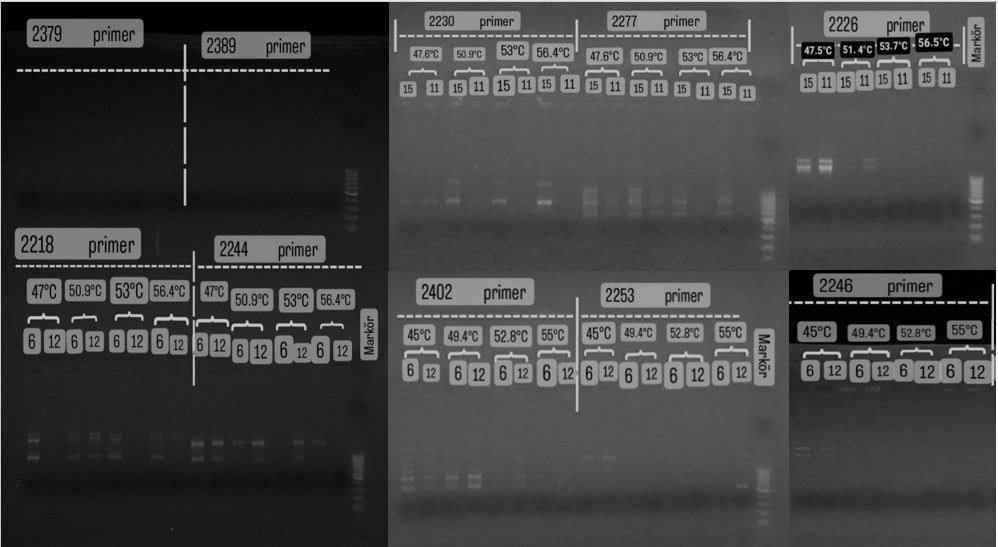
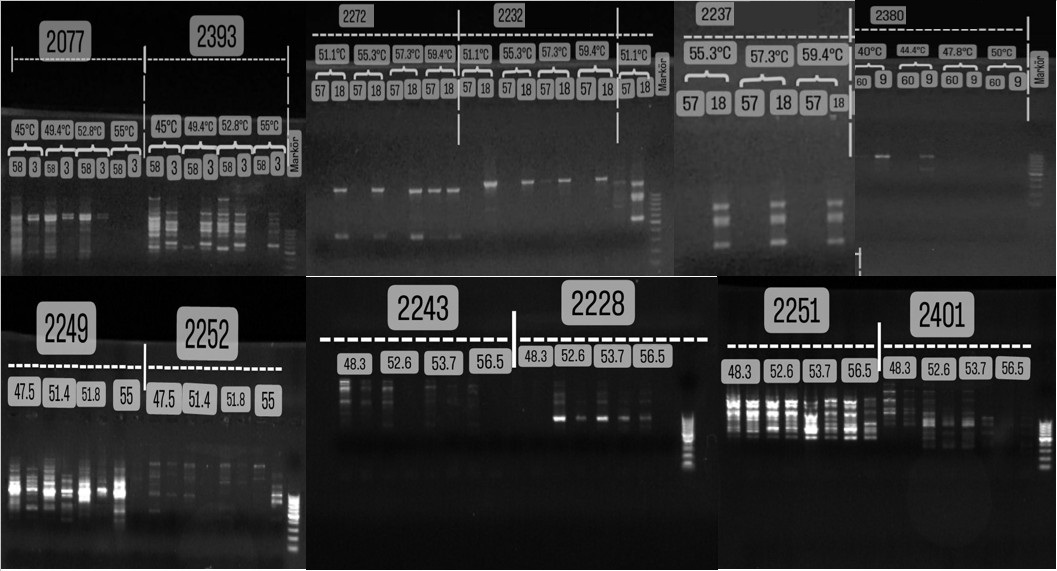
Figure (1): Agarose gel electrophoresis images showing PCR band profiles produced
by 22 different iPBS primers at various optimized annealing temperatures, revealing genetic diversity in DNA samples.
Discussion
The primary objective of this study was
to optimize Polymerase Chain Reaction (PCR) conditions based on iPBS molecular markers to accurately
and reliably determine the genetic
diversity of local sesame (Sesamum indicum L.) lines. PCR success is directly dependent on the specificity of the primers used and
the reaction parameters, particularly temperature. Therefore, PCR optimization is a critical step to ensure accurate
and efficient amplification of target DNA regions. Determining the annealing temperature
at which iPBS primers exhibit the highest polymorphism
is a crucial step in determining
optimal PCR conditions. The
results clearly demonstrate that different iPBS primers have different
annealing temperatures. This supports the
notion that each primer requires a unique optimum temperature to effectively anneal to a specific
target region, as emphasized in similar studies such as Al-Obaidi (2015), Erkol et al. (2024) and Mahyar and
Fatemeh (2021). The fact that 20 of the 22 primers used in our study
produced polymorphic bands demonstrates the high efficiency
of iPBS markers in sesame genetic studies. This high
efficiency stems from the fact
that iPBS markers target repetitive retrotransposon regions in the plant genome. Determining
the optimal annealing temperatures using the gradient PCR method increased amplification efficiency and specificity. Primer 2277 exhibited the highest
activity at 47.6°C and
50.9°C, confirming that the primer binds best to its
target at these two temperatures. In contrast, primers 2379 and 2389 yielded weak or no
amplification products, suggesting that these primers lack
suitable binding sequences for the
sesame genome or that their
annealing temperatures are still outside
the optimal range. This optimization study provides a sensitive and reliable
molecular basis for the reliable
assessment of sesame genetic diversity and the classification
of landraces. This molecular basis will make an important
contribution to future sesame breeding
programs.
Conclusion
This study demonstrates
the effectiveness of PCR analyses based on iPBS markers in determining genetic diversity in sesame genotypes. Determining optimal annealing temperatures using gradient PCR has increased amplification efficiency and specificity. The results obtained indicate that iPBS
primers can be used to assess genetic
diversity in sesame and classify local
lines. This optimization provides an important molecular basis that will
contribute to sesame breeding programs.
Author Contributions
S.E. and B.P.:
Conceptualization, Methodology,
Writing, Formal Analysis, Visualization
F.K., A.T., M.Y.: Formal Analysis, Investigation, Supervision, Writing – Review and Editing.
Funding Statement
No funding was received for
this study.
Conflicts of Interest / Competing Interests
The authors declare no competing interests.
References
Al-Obaidi, Z. (2015). Optimisation of
standard PCR programme. Kerbala Journal of Pharmaceutical Sciences, 10(1),
42–57. https://iasj.rdd.edu.iq/journals/journal/issue/14024
Doyle, J. J., & Doyle, J. L. (1990). A
rapid total DNA preparation procedure for fresh plant tissue. Focus, 12,
13–15. https://api.semanticscholar.org/CorpusID:90972282
Erkoç, K., Işık, M. İ., Erkol, Ş., Aras,
T., & Şakiroğlu, M. (2024, December 03-04). Onobrychis cinsinin genetik
varyasyonunun tanımlanması için primer arası bağlanma yerine (iPBS
işaretleyicileri) dayalı PCR analizinin optimizasyonu [Optimisation of PCR
analysis based on interprimer binding sites (iPBS markers) for the
characterisation of genetic variation in the genus Onobrychis]. 3rd
International Conference on Recent Academic Studies, Konya, Türkiye. https://as-proceeding.com/index.php/icras/3rdicras2024
Erkol, Ş., Işık, M. İ., Armağan, M., &
Aras, T. (2024, May 16-17). SCoT DNA markörleri ile Gundelia L.'nin genetik
çeşitliliğinin belirlenmesi için PCR analizinin optimizasyonu [Optimisation
of PCR analysis for determining the genetic diversity of Gundelia L. using SCoT
DNA markers]. 3rd International Conference on Engineering, Natural and Social
Sciences, Konya, Türkiye. https://www.icensos.com/
Hadeithi, Z. S. M., & Jasim, S. A.
(2021). Study of plant genetic variation through molecular markers: An
overview. Journal of Pharmaceutical Research International, 33(45B),
464–473. https://doi.org/10.9734/jpri/2021/v33i45B32828
Lakhanpaul, S., Singh, V., Kumar, S.,
Bhardwaj, D., & Bhat, K. V. (2012). Sesame: Overcoming the abiotic stresses
in the queen of oilseed crops. In N. Tuteja, S. S. Gill, A. F. Tiburcio, &
R. Tuteja (Eds.), Improving crop resistance to abiotic stress (pp.
1251–1283). Wiley-VCH Verlag GmbH & Co. KGaA. https://doi.org/10.1002/9783527632930.ch48
Mahyar, Z., & Fatemeh, N. (2021,
February 16-19). SCoT-PCR optimization in Halocnemum strobilaceum L.
[Poster presentation]. 21st National & 9th International Conference of
Biology, Semnan University, Semnan, Iran.
Özer, G., Sameeullah, M., Bayraktar, H.,
& Göre, M. (2017). Genetic diversity among phytopathogenic Sclerotiniaceae,
based on retrotransposon molecular markers. Phytopathologia Mediterranea, 56(2),
251–258. https://doi.org/10.14601/Phytopathol_Mediterr-20379
Saxena, K., & Bisen, R. (2017). Use of
RAPD marker for the assessment of genetic diversity of sesame (Sesamum
indicum L.) varieties. International Journal of Current Microbiology and
Applied Sciences, 6(5), 2523–2530. https://doi.org/10.20546/ijcmas.2017.605.283