Diagnosing Liver Cancer Through Amplification
of Mutational Extracellular mRNA: A Novel Approach
Ansar
Hussain1†, Musavir
Abbas2†, Mehwish
Kanwal3, Hafiz Muhammad Yasir1, Ghulam
Mustafa2, Zain-Ul-Abideen2, Ahmad
Hayat4, Muhammad
Qasim4, Yousaf
Raza2, Muhammad
Bilal2, Wasim
Shah2
1Chongqing Precision Medical Industry
Technology Research Institute, 400000 Chongqing, China.
2Anhui Province Biomedical Sciences and Health Laboratory, First
Affiliated Hospital of USTC, Hefei National Laboratory for Physical Sciences at
Microscale, the CAS Key Laboratory of Innate Immunity and Chronic Disease,
School of Basic Medical Sciences, Division of Life Sciences and Medicine,
University of Science and Technology of China, Hefei 230027, China; Division of
Reproduction and Genetics.
3Department of Horticulture and Pomology Fruit Quality & Storage of Horticulture
crops lab, Anhui Agriculture University, Hefei China.
4Department of Zoology,
The Islamia University of Bahawalpur, Punjab, Pakistan.
Corresponding
authors: Wasim
Shah: shah86@ustc.edu.cn and Ansar
Hussain Hussainustc@mail.ustc.edu.cn
|
Received: 02-03-2025 |
Accepted: 09-03-2025 |
First
published online:16-03-2025 |
|
|
Key words: Liquid
biopsies, Hepatocellular carcinoma (HCC), Extracellular vesicles
(EVs), SCOPE platform, Biomarkers, Early cancer detection, EV-based mRNA
profiling. |
Abstract Hepatocellular carcinoma
(HCC) remains the predominant cause of cancer-related mortality. Traditional
diagnostic methodologies are invasive and exhibit limited sensitivity for
early detection. Non-invasive alternatives, particularly liquid biopsies
utilizing extracellular vesicles (EVs), have
emerged as promising approaches. EVs contain crucial biomarkers, including
mRNA, proteins, and nucleic acids. However, the limited abundance of EV mRNA
in liquid biopsies has impeded its clinical application. To address this
limitation, researchers have developed the Self-amplified and CRISPR-aided
Operation to Profile EVs (SCOPE) platform. This innovative system integrates
CRISPR-Cas13 for RNA target identification with replication and signal
amplification, achieving subattomolar detection
sensitivity. SCOPE offers high specificity with single-nucleotide resolution
in a single-step assay. Investigators have validated probes targeting key
mutations in KRAS, BRAF, EGFR, and IDH1 genes and developed an automated
device for multi-sample analysis. SCOPE has demonstrated efficacy in
identifying early-stage lung cancer in animal models, monitoring tumor
mutational burden in colorectal cancer, and classifying glioblastoma
patients. In HCC, EV mRNA exhibits potential for non-invasive detection of
recurrence and monitoring disease progression. Current studies indicate that
EV-based mRNA profiling holds significant promise for early detection,
treatment monitoring, and recurrence prediction in HCC, offering valuable
clinical applications. The integration of advanced platforms such as SCOPE
with EV analysis could transform liquid biopsies in oncology, providing a
rapid, highly sensitive, and non-invasive method for cancer detection and
management. |
||
Introduction
Hepatocellular
carcinoma (HCC) is the most prevalent form of liver cancer, the sixth most
frequently diagnosed cancer, and the third leading cause of cancer-related
deaths worldwide (Ferlay et al., 2019).
The early detection of HCC, surveillance status, and curative treatment are
associated with significant improvements in patients' overall survival (OS) (Kim & Han, 2012). However, the incidence rate is increasing annually, and its early
diagnosis and accurate staging remain challenging (Prince et al., 2020). An additional challenge is HCC risk assessment and prevention of
cancer recurrence, along with monitoring the patients' postoperative status and
treatment response (Singal et al., 2014). Approximately 70–90% of all HCC cases develop due to liver cirrhosis,
which, in turn, can be caused by inflammation associated with hepatitis B virus
(HBV) or hepatitis C virus (HCV), exposure to toxins such as alcohol abuse and
aflatoxin B1 (AFB1), congenital disorders, and metabolic syndrome (Llovet et al., 2021). Since a large proportion of patients with HCC also have cirrhosis, it
is considered an important factor in liver injury, which leads to liver cancer.
Therefore, the discovery of minimally invasive biomarkers that can enable
precise HCC risk prediction and differentiation of HCC from non-HCC diseases is
crucial for identifying the early stages of HCC (Moldogazieva et al., 2021). Typically, tissue biopsies are invasive, and for some anatomical
sites, they are not easily obtainable. They also provide a limited
representation of intratumoral and intermetastatic genetic heterogeneity because tumors are
heterogeneous entities containing various subpopulations of cells that feature
different lesions (Ignatiadis
et al., 2021; Martins et al., 2021). Furthermore, cancer cells undergo genetic and epigenetic changes over
time and can evolve dynamically, guided by microenvironmental stimuli and
clonal selection due to therapeutic pressure. This results in further tumor
heterogeneity (Martins et al., 2021), thus affecting the accuracy of the examination and the therapeutic
decisions made based on it. Additionally, surgical biopsies have limitations in
terms of time, repeatability, patient age, and cost and can occasionally cause
harmful clinical complications (Martins et al.,
2021).
Consequently, they are not suitable for highlighting the overall tumor profile,
identifying lesions in different locations, or longitudinal monitoring of the
disease.
Identifying genomic variability in liquid
biopsies can significantly advance precision oncology
Liquid biopsies are emerging as key tools for addressing
challenges in the prognosis, diagnosis, and monitoring of disease progression.
The SCOPE technique offers several advantages, including reduced invasiveness,
lower cost, and the ability to provide up-to-date information on tumor status.
In some cases, it can also address the problem of tumor heterogeneity or
multiple metastatic changes (Killingsworth et al., 2021). These biopsies involve examination of bodily fluids, primarily blood,
but can also include urine, saliva, cerebrospinal fluid (CSF), and bone marrow (Pantel & Alix-Panabières, 2019). In precision oncology, liquid
biopsies enable the collection of dynamic molecular data regarding the entire
tumor through minimally invasive and repeatable tests (Ignatiadis et al., 2021; Killingsworth et
al., 2021).
Consequently, both scientists and medical professionals use liquid biopsies to
track tumor evolution and heterogeneity (Parikh et al., 2019). The real-time insights gained from these biopsies can influence
patient care in various ways, such as early detection of lesions, tracking of
minimal residual disease, guiding personalized treatment decisions based on
resistance profiles, and monitoring tumor recurrence (Parikh et al.,
2019).
Liquid
biopsy components, including circulating tumor DNA (ctDNA),
circulating tumor cells (CTCs), and exosomes, reflect the phenotypic and
genotypic properties of tumor cells. Tumor-derived exosomes transport a diverse
array of molecular cargoes, such as microRNAs, long non-coding RNAs, and
circular RNAs, which are subsequently delivered to recipient cells. Exosome
biogenesis involves invagination of the plasma membrane to form early
endosomes, which subsequently mature into late endosomes or multivesicular
bodies (MVBs). MVBs either fuse with lysosomes for degradation or merge with
the cell membrane to release exosomes into the extracellular milieu.
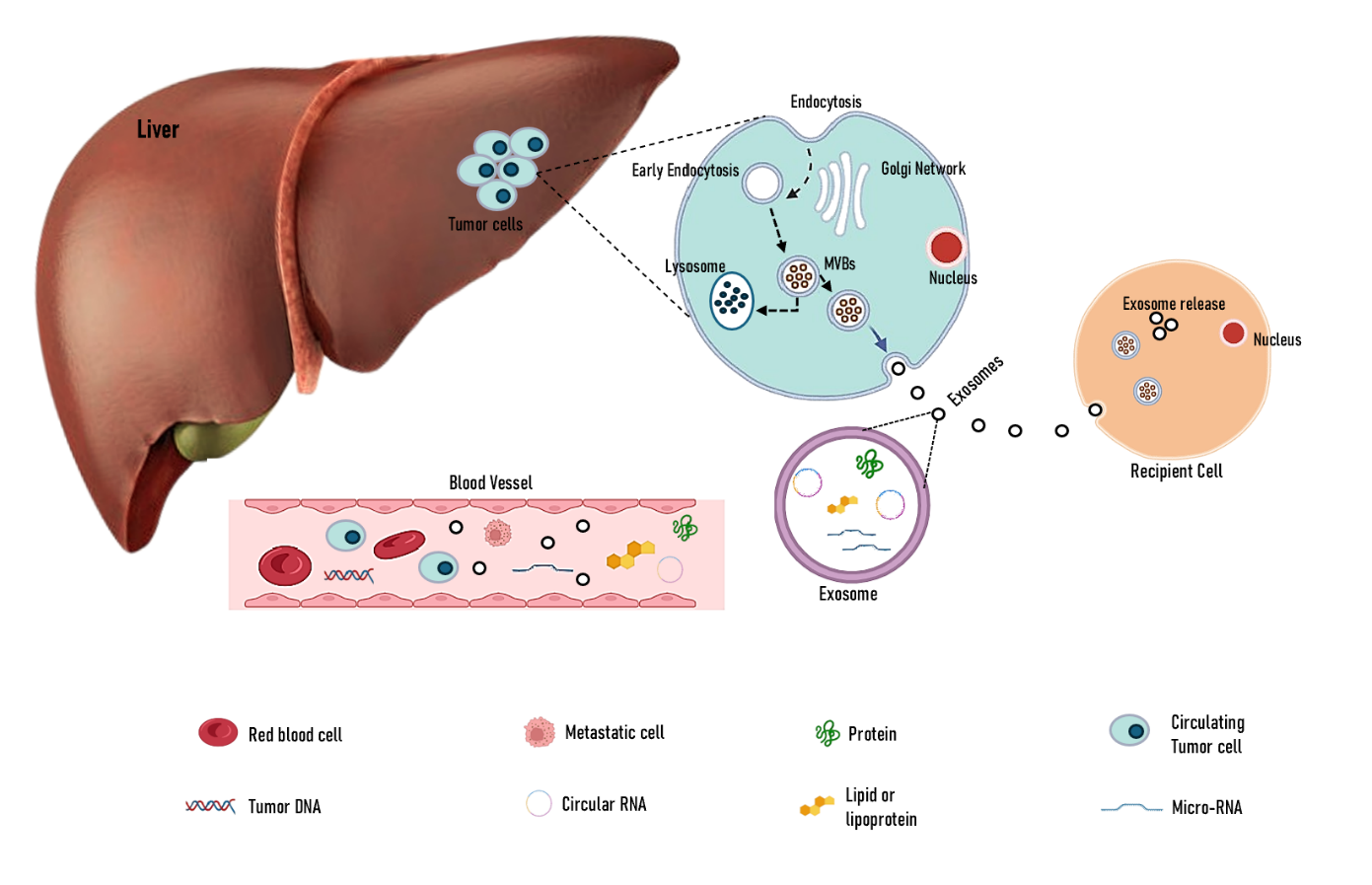
Figure 1. Overview of Liquid Biopsy Components and Exosome Biogenesis in
Liver Tumor Cells.
Extracellular
vesicles (EVs) constitute a heterogeneous group of
membrane-bound structures that are secreted by all cell types into various
biological fluids. Encapsulated within a phospholipid bilayer, EVs contain a
diverse array of bioactive molecules, including proteins, lipids, and nucleic
acids, originating from their parent cells (Ferlay et al.,
2019; Kim & Han, 2012).
Based on their dimensions, biogenesis mechanisms, and molecular composition,
EVs are broadly classified into two primary categories: exosomes and
microparticles (MPs). These distinctions are essential for elucidating the
multifaceted functions (Prince et al.,
2020).
Under physiological
conditions, EVs are continuously secreted. However, their release is
significantly elevated in pathological states such as inflammation and cancer (Martins et al.,
2021). Upon release, EVs can interact with recipient cells through processes
such as endocytosis, delivery of functional cargo, and eliciting a diverse
array of cellular responses, as explained in Figure 1 (Ignatiadis et al.,
2021; Llovet et al.,
2021). This capability enables EVs to influence both immunostimulatory and
immunoinhibitory pathways, with their effects varying depending on the
originating cell type and the specific bioactive content they carry (Singal et al.,
2014).
EVs
play a crucial role in numerous biological processes, including inflammation,
immune signaling, coagulation, vascular reactivity, angiogenesis, and tissue
repair (Zarà et al.,
2019). They function as essential mediators of intercellular communication,
facilitating the transfer of molecular signals between cells (Everaert, 2020).
These vesicles transport vascular growth factors, such as VEGF and ANGPT2, to
endothelial cells (ECs), modulating their biological characteristics and
promoting angiogenesis via pathways such as AKT/eNOS.
Hypoxia further enhances EV-mediated angiogenic signaling, with elevated exosomal miR-155 levels correlating with VEGF expression
and vascular density. In addition to ECs, macrophages contribute to
angiogenesis through EV-transferred miRNAs, which modulate
epithelial-mesenchymal transition (EMT) and vascular permeability. Furthermore,
EVs can function as platforms for enzymatic activity, further expanding their
range of physiological functions (Block et al.,
2022). These distinctive characteristics have led to their recognition as
significant factors in health and disease, particularly in the context of liver
disease, where they form complex regulatory networks with hepatic macrophages (Cheng et al.,
2024).
In
pathological conditions, EVs have garnered considerable attention owing to
their involvement in various disease mechanisms, including autoimmune
disorders. They are being increasingly investigated as potential biomarkers for
cell activity or death, offering insights into disease progression. Moreover,
EVs show promise as innovative drug delivery vehicles, leveraging their
inherent ability to transport molecular cargo across biological barriers (Makrygianni & Chrousos, 2023).
EV cargo comprises proteins (e.g., heat shock proteins, adhesion molecules, and
growth factors), nucleic acids (notably RNA, including miRNA, mRNA, and ncRNA),
and lipids (e.g., ceramides and sphingolipids), which influence recipient cells
and serve as potential biomarkers. Proteins such as CD63 and CD81 are
particularly significant given their roles in EV formation and cargo sorting (Cheng et al.,
2024).
The emerging understanding of EVs underscores
their importance in both physiological and pathological contexts, thereby
opening new avenues for research and therapeutic development (de Lima et al.,
2020; Singal et al.,
2014). Their multifaceted roles in disease pathogenesis, cellular
communication, and potential clinical applications have rendered them a focus
of increasing scientific and medical interest.
Techniques for Isolating Extracellular Vesicles
(EVs)
The pre-analytical phase
Successful
outcomes in EVs isolation commence with the initial steps of blood extraction
and acquisition of bodily fluids. The generation of artifactual EVs becomes
more probable during experimental conditions due to multiple factors that
particularly affect platelet-derived or red blood cell-derived EVs owing to
their high sensitivity. The methodology of blood collection, the specifications
of tubes and anticoagulants, transportation protocols, and the duration between
sample collection and testing contribute to EVs isolation (Lacroix et al.,
2012). The development of standardized procedures is ongoing, which will
mitigate pre-analytical variables (Witwer et al.,
2013). The isolation process for EVs should be conducted within the first 2 hr after sample collection, during which the samples should
undergo minimal movement. Exosome isolation initiates with centrifugation at
low speeds, followed by filtration or size-exclusion chromatography (SEC) before
high-speed centrifugation at 100,000 g sediments them (Witwer et al.,
2013).
Ultracentrifugation of density gradient is an efficacious method to enhance
purity. The recommendations by ISTH 2010 and Lacroix et al., 2012 indicate that MPs should be isolated through double
centrifugation of whole blood at 2500 g for 15 minutes at room temperature.
The Analytical phase
The
analysis of EVs is complicated due to their submicron dimensions and
heterogeneous characteristics regarding origin, size, and composition. A standardized
method to characterize EVs does not currently exist. Flow cytometry has
remained the primary technique for EVs characterization for more than two
decades due to its ability to analyze EVs composition along with their quantity
(György et al.,
2011; Horstman et al., 2004; Edwin
Van der Pol et al., 2012).
The conventional flow cytometry method utilizing light scattering detection
enables the detection of only large EVs that measure approximately 1 μm (Arraud et al.,
2014; Chandler et al., 2011; E Van
Der Pol et al., 2012).
The detection of the majority of EVs, except the smallest ones, is feasible
through a recent modification of flow cytometry that uses fluorescence
intensity-based detection techniques (Arraud et al.,
2015; Van Der Vlist et al., 2012).
Improved flow cytometry instrumentation will advance
research by enabling the detection of smaller objects. Evolutionary progress in
EVs science began with platelet MPs detection by electron microscopy, which
provided accurate images of EVs and fundamental information about size and
phenotype expression (Aalberts et
al., 2012; György et al., 2011;
Heijnen et al., 1999; Wolf, 1967).
Unprecedented views of EVs in blood plasma and other body fluids have been
achieved through recent applications of cryo-electron microscopy, a method that
optimally preserves complex objects (Arraud et al.,
2014; Zonneveld et al., 2014). Scientists
established a comprehensive characterization of pure plasma EVs through their
work, which demonstrated spherical EVs between 50-500 nm as the predominant
population, while plasma also contains larger tubular EVs and membrane
fragments exceeding 500 nm (Arraud et al.,
2015).
This research found that Phosphatidylserine exposure affected approximately
fifty percent of all detected EVs. Cryo-electron microscopy surpasses other
techniques for revealing native biological fluid content at nanometer
resolution by demonstrating the presence and variety of individual objects along
with micrometer-sized immune complexes near EVs within arthritis patient
synovial fluids. Electron microscopy remains a time-consuming and expensive
analytical method that requires skilled personnel to operate, thus creating
immediate limitations for clinical application. Western blotting and ELISA
serve as commonly used methods for EVs phenotype examination by measuring intravesicular or membrane protein markers through antibody
detection according to (Revenfeld et
al., 2014).
EVs were examined by conducting RT-qPCR to reveal their RNA content.
High-throughput EVs analysis has expanded through the introduction of two
recent technologies: nanoparticle tracking analysis (NTA) and tunable resistive
pulse sensing (TRPS). These techniques excel at creating size-related
observations for small particles (50 nm) to facilitate the identification of
purified exosome preparations. These two methods prove challenging to use for
heterogeneous samples such as pure plasma since they lack the ability to
distinguish EVs from contaminants such as lipoproteins (Zarà et al.,
2019).
New performance advancements and improved fluorescent dyes within flow
cytometers, along with new technological developments, will enhance EVs
analysis capabilities.
Detecting of EVs mRNA
revealed key somatic driver mutations essential for tumor initiation and
growth.
Extracellular
vesicles are emerging as significant targets in liquid biopsy research (Jo et al.,
2023; Shao et al., 2018). These minute particles, less than 1 µm in diameter, transport various
molecular components, including nucleic acids, proteins, and metabolites,
effectively acting as cellular proxies (Dixson et al.,
2023).
Analysis of EVs messenger RNA (mRNA) can yield valuable clinical insights (Nomura et al.,
2009). Extracellular vesicles mRNA can indicate the
presence of somatic driver mutations such as KRASG12D and BRAFV600E, which are
critical for tumor development (Skog et al.,
2008).
Additionally, while EVs rarely contain nuclear proteins associated with drug
resistance, they carry the corresponding mRNA, providing information about the
resistance status (Daane et al.,
2022; van de Haar et al., 2023).
Vesicular encapsulation of EV mRNAs shields them from nucleases in biofluids,
enabling the extraction of intact, high-quality nucleic acids (Park et al.,
2021).
These characteristics make EVs a promising source of nucleic acids,
complementing the advantages of circulating tumor DNA (ctDNA).
However, technical constraints have limited the clinical application of EVs.
Most EVs RNA is non-coding, and the mRNA levels in EV samples can be extremely
low. For example, even abundant mRNA species, such as GAPDH, are detected at a
rate of only one copy per 104–106 EVs, in contrast to microRNAs at one copy per
102 EVs (Noerholm et
al., 2012; Wei et al., 2017).
This
disparity has led most proof-of-concept studies to concentrate on miRNA
detection (Park et al.,
2021; van de Haar et al., 2023; Wei et al., 2017).
Furthermore, tumor-derived EVs comprise a small fraction (<5%) of total
circulating EVs (Noerholm et
al., 2012).
The scarcity of EV mRNAs and low abundance of tumor-derived EVs necessitate
large sample volumes (exceeding 2 ml of plasma) and advanced technologies such
as droplet-digital polymerase chain reaction (PCR) and next-generation
sequencing. This reduces the competitive edge of EV tests and hinders their
incorporation into the standard preclinical and clinical assays.
A novel EVs mRNA test inspired by CRISPR
technology was developed. CRISPR systems are increasingly utilized in molecular
diagnostics due to their sequence-specific nuclease activity (Kaminski et
al., 2021; Pickar-Oliver & Gersbach, 2019). CRISPR-associated (Cas) proteins function as endonucleases when they
recognize target nucleic acids (Abudayyeh & Gootenberg, 2021). This property has been exploited to amplify signals through the
indiscriminate cleavage of reporter probes, such as single-stranded DNAs tagged
with fluorescent dye and quencher pairs. However, applying CRISPR assays to EV
mRNA is challenging due to the low abundance of targets, often requiring
pre-amplification to replicate mRNA and enhance assay kinetics. This step can
introduce replication errors and biases, potentially leading to misleading
results (Kebschull & Zador, 2015; Potapov & Ong, 2017). To address this limitation, Cas activity was repurposed to directly
recognize and replicate the target mRNA in situ, eliminating the need for
pre-amplification and its associated errors. This innovative approach ensures
high analytical sensitivity while maintaining sequence specificity, enabling
precise detection of low-abundance mRNA targets (Song et al.,
2024).
This review aimed to elucidate the clinical significance of the SCOPE technique
in liquid biopsies for hepatocellular carcinoma (HCC) and examine its potential
for prognosis, diagnosis, and monitoring of cancer progression. The mRNA
amplification method and its clinical applications will also be analyzed, with
a particular emphasis on the development and future prospects of the SCOPE
technique.
SCOPE: A CRISPR-enhanced platform for EV mRNA
detection
The SCOPE (Self-amplified and CRISPR-aided Operation to
Profile EVs) platform is an innovative integrated assay for accurate EV mRNA
detection and monitoring. It merges the Cas13a machinery with novel signaling
templates, enhancing both specificity and sensitivity. SCOPE operates by Cas13a
recognizing target RNA sequences, triggering a dual amplification process that
boosts both RNA targets and fluorescent signals, thereby ensuring robust
detection.
SCOPE's exceptional
selectivity of SCOPE, attributed to Cas13a, allows precise single-nucleotide
polymorphism differentiation. The platform achieved high sensitivity and
detected subattomolar concentrations through its dual
amplification mechanism, which was further refined by systematic optimization.
SCOPE's versatility of SCOPE has been demonstrated in various applications,
including early-stage lung cancer detection in animal models during preclinical
studies. In clinical settings, it has been used to track cancer mutational
burdens in patients with colorectal cancer (CRC) undergoing standard
treatments, providing valuable insights into disease progression and treatment
responses. Moreover, SCOPE has effectively identified crucial mutations in
glioblastoma multiforme (GBM), facilitating patient stratification for more
targeted treatment approaches (Song et al.,
2024).
The
implementation of this advanced assay offers significant opportunities in both
preclinical and clinical settings. It serves as a tool for understanding cancer
progression mechanisms, identifying the emergence of drug resistance, and
assessing tumor responses to various therapies. Beyond its scientific
applications, SCOPE has the potential to significantly impact clinical
workflows and drug trial decision-making processes. By accelerating standard
decision making in clinical trials and enhancing the utility of extracellular vesicles (EVs) in liquid biopsy applications, SCOPE bridges
the gap between cutting-edge molecular diagnostics and practical clinical use.
In
summary, the SCOPE platform introduces a novel approach for incorporating
extracellular vesicle profiling in cancer research and clinical practice. Its
ability to provide highly accurate, sensitive, and actionable insights into
tumor biology makes it an invaluable tool for advancing oncology and
personalized medicine.
SCOPE technology
setup
The
SCOPE technology platform integrates several cutting-edge components to boost
the effectiveness of EV mRNA detection (Song et al.,
2024). A key aspect of this approach is the use of polymer-coated tubes (pDMAEA-coated PCR tubes) for rapid and efficient nucleic
acid extraction. When an aqueous sample is introduced, the positively charged
polymer binds to negatively charged nucleic acids, forming polyplexes that are
then isolated through centrifugation. The SCOPE reaction occurred directly in
the same tube, eliminating the need for sample transfer and streamlining the
process. The system includes a fluorescence detection device with a tray-type
heating block, a fluorescent optical detector, and a line scanner capable of
analyzing up to 16 samples simultaneously. Separate fluorescent
excitation/detection headers allow for one- or two-color measurements with high
consistency and uniform temperature regulation, maintaining variations within
0.5°C across samples. Designed for ease of use, the system utilizes standard
lab equipment, such as thermal cyclers, and requires minimal sample volumes
(e.g., EV isolates from less than 100 μL of plasma).
An intuitive graphical interface controls this process and supports its
applications in clinical and preclinical molecular diagnostics.
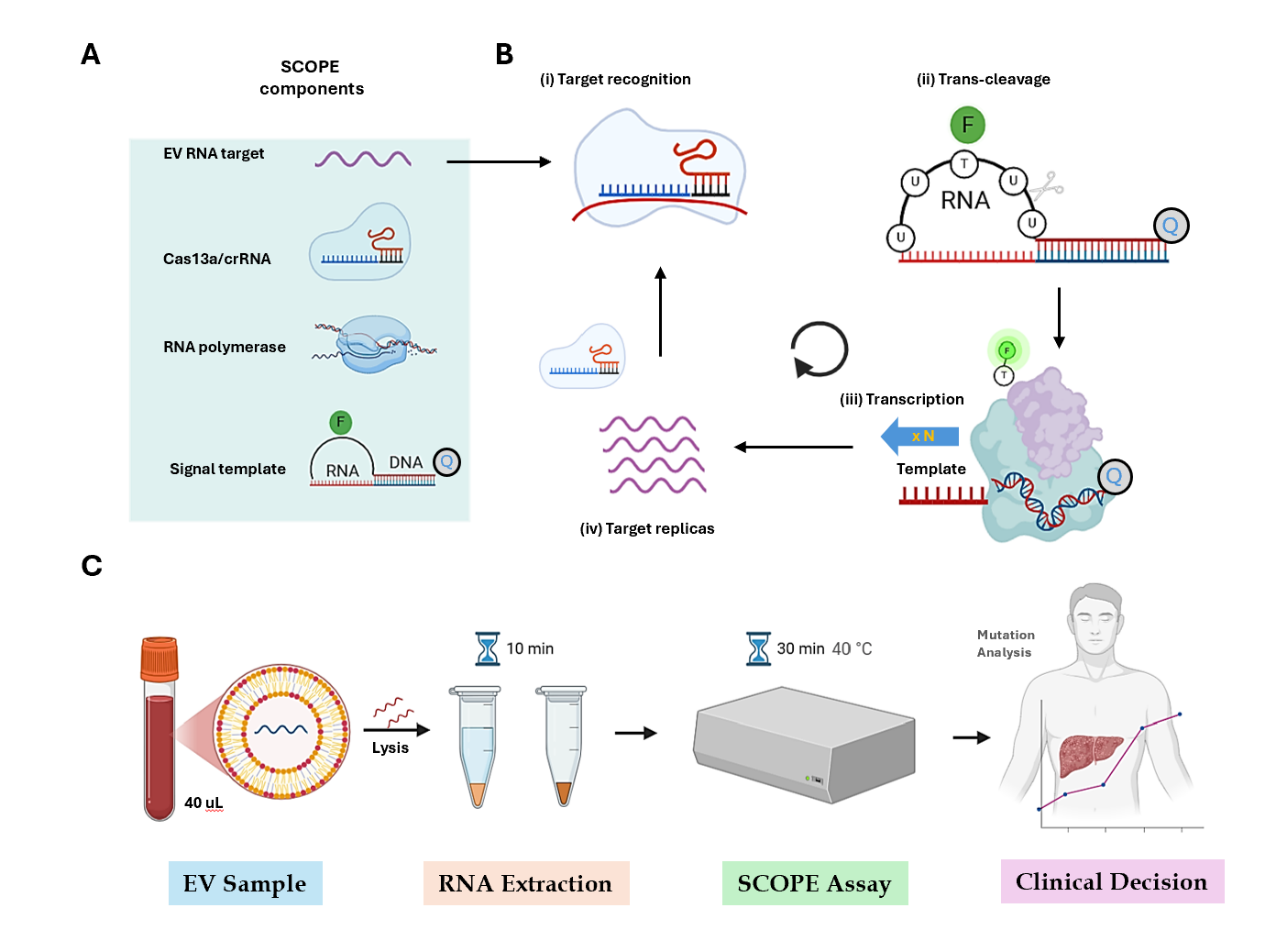
Figure
2: SCOPE workflow for on-site diagnostics. (A) Illustrating diverse elements
of SCOPE, such as EV RNA target, Cas13a, RNA polymerase, and signal template.
(B) Combined SCOPE components identify the target for replication. (C) The
SCOPE system enables rapid on-site molecular testing. Initially, EVs were
extracted from the clinical specimens and broken down. The resulting extracts
were placed in specially treated containers for RNA isolation, which required
approximately 10 min. Subsequently, the SCOPE reaction was performed in a small
transportable device for 30 min. In total, the test delivers molecular data
within an hour, allowing same-day clinical choices to be made.
SCOPE working
principle
The SCOPE operational mechanism integrates CRISPR-Cas13a
recognition with RNA amplification. The process begins with a combination of
Cas13a, CRISPR RNA (crRNA), T7 polymerase, signal template, and
deoxyribonucleotide triphosphates. The signal template, composed of RNA and
DNA, plays a vital role in linking the Cas13a/crRNA and T7 polymerase
reactions. Initially, Cas13a/crRNA attaches to the target RNA, thereby
activating its ribonuclease function. This action cleaves the RNA portion of
the signal template, releasing fluorescent dye molecules, and generating a
detectable signal. Simultaneously, T7 RNA polymerase multiplies the target RNA,
producing numerous copies that are then recognized and cleaved by Cas13a/crRNA,
creating a powerful amplification cycle. SCOPE achieved high specificity by
incorporating synthetic mismatches in crRNA, allowing Cas13a to distinguish
between RNA sequences, even at the single-nucleotide level. Additionally, dual
amplification through Cas13a cleavage and RNA replication ensures high
sensitivity. The isothermal reaction occurs at 40°C, enabling the completion of
the entire process in under an hour within a single tube (Vitale et al.,
2021). These features make SCOPE efficient, sensitive, and appropriate for
routine laboratory use, with a significant potential for clinical diagnostic
applications (Figure 2).
Kinetics of the SCOPE Assay
Researchers have employed analytical modeling to
thoroughly examine the kinetics of the SCOPE assay, shedding light on the
underlying mechanisms and reaction dynamics (Song et al.,
2024). The SCOPE method combines two separate catalytic processes facilitated
by a signal template, thereby enabling efficient RNA detection and
amplification. The initial process involves the binding of the Cas13a/crRNA
complex to the target RNA, triggering fluorescent signaling through the cleavage
of the RNA segment within the signal template. The second process uses RNA
polymerase to replicate the target RNA using a DNA sequence incorporated into
the signal template. When studied independently, these catalytic activities
produced linear increases in the reaction products over time, aligning with
zeroth-order reaction kinetics under specific assay conditions (Song et al.,
2024).
However, coupling these processes via the signal template significantly alters
the reaction kinetics, approximating a first-order reaction rate and
substantially enhancing the efficiency. The SCOPE signals exhibited exponential
growth, reaching a plateau within 30 min.
A key feature of the SCOPE assay is its ability to
suppress off-target RNA amplification. The amplification process requires the
Cas13a/crRNA complex to first recognize and bind to the target RNA, activating
Cas13a to cleave the RNA segment in the signal template and initiate RNA
polymerase activity. If the RNA segment remains intact, polymerase activity is
inhibited, likely because the intact loop configuration of the signal template
physically obstructs the polymerase from accessing the promoter region and
initiating transcription. To enhance the performance of the SCOPE assay,
researchers have refined the signal template design and optimized the reaction
conditions for maximum signal intensity (Figure 3A). The validation experiments
(Figure 3B) demonstrated that the optimal analytical signal was achieved only
when all crucial assay components—target RNA, Cas13a, crRNA, and T7 RNA
polymerase—were present (Song et al.,
2024).
Excluding RNA polymerase from the reaction significantly reduced the signal
intensity by halting additional RNA target generation. These findings highlight
the robustness and efficiency of the SCOPE assay and confirm its reliability
for RNA detection and amplification. The ability of this assay to combine rapid
signal generation with high specificity makes it a valuable tool for various
molecular diagnostic applications.
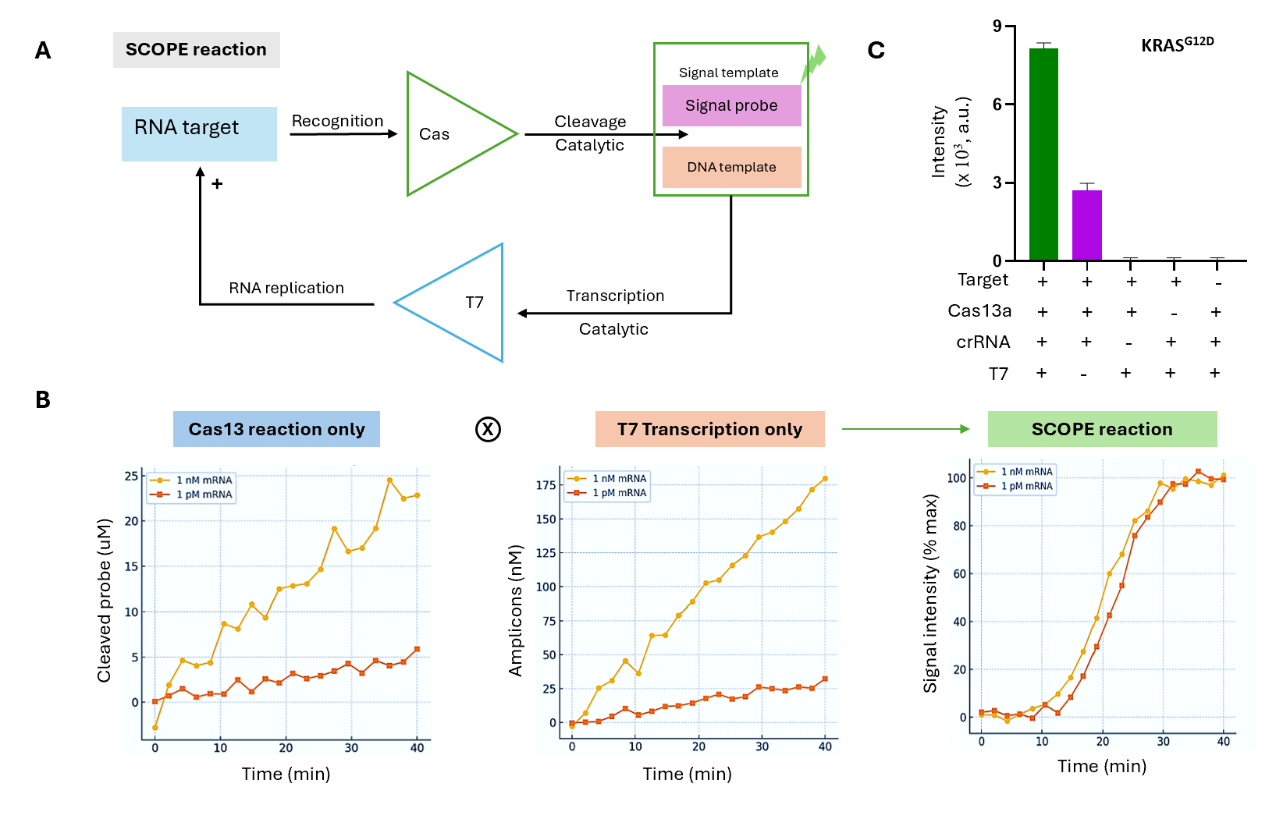
Figure 3: SCOPE assay dynamics. (A) The SCOPE method integrates two enzymatic processes: Cas13a/crRNA
produces fluorescence by breaking down RNA in the signal template, whereas T7
polymerase multiplies RNA targets. (B) When operating independently,
Cas13a/crRNA and T7 reactions displayed linear product increases over time.
However, when combined with SCOPE, they result in exponential signal
enhancement, achieving a plateau within 30 min. Experimental confirmation
demonstrated a peak signal when all components were present; eliminating T7
polymerase or impeding mRNA recognition diminished signal strength. (C) The starting
KRASG12D RNA concentration was 1 nm.
Conclusion and Future Prospects
Self-amplified and CRISPR-aided Operation to Profile EVs
(SCOPE) technology represents a significant advancement in liquid biopsy
applications and offers exceptional sensitivity, specificity, and
accessibility. Its capacity to detect genetic mutations at subattomolar
concentrations and differentiate single-nucleotide variations surpasses
conventional diagnostic tools. By facilitating the analysis of EV mRNA, SCOPE
addresses critical challenges in early cancer detection and monitoring,
particularly in malignancies such as liver cancer, where existing diagnostic
approaches often fail to identify the disease at an early stage. Furthermore,
the versatility of this technology extends to monitoring treatment responses in
real time, assessing minimal residual disease, and tumor subtyping. With its
low cost (less than $4 per marker), rapid assay times (approximately 30
minutes), and compatibility with standard laboratory equipment, SCOPE has the
potential to revolutionize clinical and research practices by rendering liquid
biopsy both accessible and practical for routine use (Daane et al.,
2022).
While SCOPE shows significant promise, several obstacles
must be overcome to fully harness its potential. One major issue is the
unintended isolation of other extracellular RNA carriers, including
lipoproteins and EVs, from platelets during the sample preparation process
using size exclusion chromatography (SEC). These unwanted components can
increase background noise, make data analysis more complex, and potentially
decrease the diagnostic precision. To tackle this problem, advanced techniques,
such as single-vesicle imaging, can be employed. This method involves labelling
and monitoring specific proteins on vesicle surfaces, which can greatly improve
the accuracy of target identification and enhance the reliability of SCOPE
diagnostic results.
Future research should focus on validating the SCOPE
across a broader spectrum of cancers and diverse treatment settings.
Integrating the analysis of EV mRNA with circulating tumor DNA (ctDNA) could provide a more comprehensive molecular tumor
profile by capturing both transcript-level changes and unique genomic
alterations, such as promoter mutations and methylation patterns (Everaert, 2020).
This dual approach would bridge the gap between transcriptomics and genomics,
ensuring improved diagnostic precision and enabling a deeper understanding of
tumor biology. Furthermore, optimizing SCOPE for multimodal treatment
strategies and establishing cancer-specific timelines for EV analysis after
surgery or therapy initiation will enhance its clinical relevance. For
instance, EV mutational loads observed in patients with colorectal cancer (CRC)
demonstrate fluctuations after surgery and during chemotherapy, highlighting
the importance of defining optimal timeframes for sample collection to refine
prognostic predictions and guide adjuvant therapy decisions (Daane et al.,
2022).
Self-amplified and CRISPR-aided Operation to Profile EVs
(SCOPE) exhibits considerable promise in preclinical drug development, offering
swift and accurate insights into tumor biology that can accelerate the
assessment of therapeutic responses. Its capacity to identify point mutations
with minimal interference, even at low variant allele frequencies (such as
0.01%), outperforms many advanced techniques including digital PCR and BEAMing PCR, making it an invaluable tool for drug testing.
These attributes can streamline the drug discovery processes, enhance
experimental therapies, and promote clinical translation.
The revolutionary aspect of SCOPE lies in its ability to
provide same-day results and enable real-time clinical decision-making. This
rapid turnaround, combined with its high sensitivity and cost-efficiency, makes
it particularly well-suited for applications such as early cancer detection,
monitoring recurrence, and personalizing treatment. When integrated with
complementary methods, such as the analysis of EVs mRNA and ctDNA,
SCOPE offers a comprehensive molecular profile of tumors, ensuring more precise
diagnoses and customized treatment plans.
Addressing current limitations, such as improving EVs
specificity, expanding the range of cancers analyzed, and exploring its utility
in multimodal treatment contexts, will be critical to realizing SCOPE's full
potential of SCOPE. For instance, in cases such as EGFRvIII
deletion, in which designing a specific ctDNA assay
is challenging, the detection of EV mRNA through SCOPE provides a more feasible
and effective alternative. Additionally, incorporating new imaging techniques
to study the origin of EVs-associated mRNA will further enhance its utility as
a diagnostic tool.
Building on these advancements, SCOPE is well-positioned
to redefine cancer diagnostics and treatment monitoring. Its ability to provide
a reliable, accessible, and comprehensive molecular analysis platform will
undoubtedly enhance precision oncology, improve patient outcomes, and establish
a new standard for liquid biopsy in both academic and clinical settings.
Data
and code accessibility
The authors confirm that
the data presented in the article and additional data can be provided by the
corresponding author upon request.
Author contributions
A.H., and M.A., have
gathered data, outlined and finalized the initial manuscript draft. M.K., G.M.,
Z.U.A., A.H., M.Q., Y.R., and M.B., helped to analyze data. W.S., provided
revisions and finalized the manuscript. All authors have reviewed and approved
the final manuscript.
Conflict
of Interest: The authors have disclosed no conflicts of
interest.
Funding:
N/A
Approval for publication: After
reviewing the manuscript, the authors have decided to submit it to the
publication. The authors declare that nothing in the study has ever been
published before or is presently being considered for publication anywhere.
References
Aalberts, M., van
Dissel-Emiliani, F. M., van Adrichem, N. P., van Wijnen, M., Wauben, M. H.,
Stout, T. A., & Stoorvogel, W. (2012). Identification of distinct
populations of prostasomes that differentially express prostate stem cell
antigen, annexin A1, and GLIPR2 in humans. Biology
of reproduction, 86(3), 82,
81-88.
Abudayyeh,
O. O., & Gootenberg, J. S. (2021). CRISPR diagnostics. Science, 372(6545),
914-915.