Tet(X)
Functionality: The Tet(X)
gene is functional in environmental (aerobic) Sphingobactrium species,
but also found in anaerobic Bacteroides species.
Answerable for
the rapid unfolding of unique factors throughout bacterial groups around the
sector. Horizontal gene transfer is associated with three primary mechanisms:
(a) Conjugation, plasmid transfer from one bacterium to every other;
(b)
transduction, viral-mediated (phage) gene switch; and (c) transformation, the
uptake of bare DNA through the cell wall, and the incorporation of that DNA
into the existing genome or plasmids (Kumarasamy et al. 2010; Levy 2002). The
Tet genes listed in desk 2.1 are related to conjugative, nonconjugative, and
mobilizable plasmids, transposons, and conjugative transposons (Fig. 2.2).
1.2.1 Intrinsic
Resistance
In a few
instances, a sort of bacteria will live on antibiotic remedy and multiply due
to the fact it's miles intrinsically resistant. For instance, even though many
kinds of bacteria have cellular walls, a few don’t. An antibiotic like
penicillin that prevents mobile-wall building can’t harm a bacterium that
doesn’t construct a cell wall within the first location (Fig. 2.3).
1.2.2 Obtained
Resistance
Bacteria also
can collect resistance. This takes place whilst a sort of bacteria adjustments
in a manner that protects it from the antibiotic. Microorganism can accumulate
resistance in ways: both via a brand-new genetic exchange that allows the
bacterium to continue to exist, or by using DNA from a bacterium is already
resistant.
1.2.3
Genetic alternate
So how can an
easy DNA alternate protect bacteria from antibiotics? Consider, DNA offers
commands to make proteins, so a change in DNA can cause a
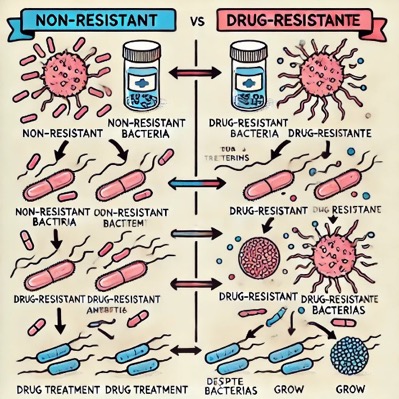
Fig. (2) Diagram displaying the distinction
among non-resistant bacteria and drug-resistant bacteria. Non-resistant
bacteria multiply, and upon drug treatment, the microorganisms die.
Drug-resistant microorganism multiply as nicely, but upon drug treatment, the bacteria
continue to spread (adapted from Wikipedia)
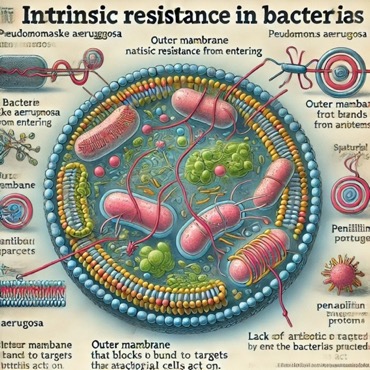
Fig. (3) Intrinsic resistance
Right
here is the diagram illustrating intrinsic resistance in microorganisms. It
suggests how certain organisms, like Pseudomonas aeruginosa, have
natural resistance mechanisms along with an outer membrane that blocks
antibiotic access and a lack of antibiotic targets. Let me know if case you'd
like any changes or additional detail
Fig.
(4) Genetic change
Right
here is the diagram illustrating genetic changes, together with mutations,
recombination, and horizontal gene switch.
Alternate in a protein.
Occasionally, this DNA exchange will influence the protein’s shape. If this
takes place in the place of the protein in what way an antibiotic act, the
medicine can furthermore not anymore be able to recognize at which point it
wishes commotion allure task.
Adaptations in this manner concede
possibility prevent a medicine from accepting into the container or hamper the
medicine from operating once it’s central. As directly as an exchange occurs,
it grants permission to spread in a people of bacteria by way of approaches
like duplication or DNA switch (Fig. 2. 4).2.2.4r DNA transfer bacterium are
superior at giving genes, which contain genes for medicine fighting. They are
intelligent to dimension fighting genes that have been in the culture, apart
from newancestral changes that stand. Either you explored Agent Antibiotic, you
proverb a germ accompanying a medicine-fighting gene presents a copy of that
deoxyribonucleic acid to another germ. This arrangement is referred to as a
lateral deoxyribonucleic acid switch. There are various habits bacteria can
transfer DNA, for instance, bacteria can receive congested accompanying in a
way bug popular as a bacteriophage. As part of its behaviors phase, the
bacteriophage bundles DNA. As long as the bacterium dwindles, those programs of
DNA (that sporadically contain medicine resistance genes) are freed and concede
the possibility stop living up and secondhand by various bacteria (Fig. 2.5)
Fig.
(5) DNA transfer
.3
Tetracycline resistance genes
Tetracyclines are individual of the
oldest instructions used medicines and the first popular elegance of medicines.
Tetracyclines communicate with accompanying bacterial ribosomes through
reversible attachment to the ribosome that blocks protein association.
Tetracyclines are energetic towards a roomy array of gram-accurate,
gram-horrific, anaerobic, and cardia microorganisms, field-obstruction-unfixed
bacteria, intercellular microorganisms and bloodthirsty flagellates.
Tetracyclines are almost cautious
and earlier compounds are modest and they were common for objective, veterinary
and ground purposes 60 years (Roberts 2005b).{17} However, for this affiliate
it will include a gram-beneficial microorganism Mycoplasma, Ureaplasma,
box-delider-free, similar to Mycobacterium, Nocardia and Streptomyces. primary
Tcr microorganisms were identified in isolates from the 1950s (Watanabe et al.
1972). bacteria get the opportunity to beautify themselves, unlike
tetracyclines, by metamorphosis, while the ripeness of microorganisms adorns
the medicines of conflicting causes that accept new ones genes that (a) pull
tetracycline out of the container (efflux); (b) secure the ribosome before
action of tetracyclines; or c) enzymatically deactivate tetracyclines (Table
2.1).
2.3.1
Discharge
The first drug-resistant efflux
proteins were classified in the 1950s in Japan regions later speculated to be
found on conjugative plasmids (Watanabe 1963). nowadays, there are professional
27 congenitally unconnected efflux genes typical of systematization drug-Hþ
electricity-powerless transmembrane series (TMS) proteins that span central
field sheath lipid bilayer nine–14 duration. Those proteins were indifferent
for seven different organizations involved, number of TMS donations (9–14), G þ
C % (guanine-cytosine) deoxyribonucleic acid and correspondence with other tet
efflux genes (Thaker et al. 2010).{18} these efflux proteins usually release
the drug and doxycycline but do not
2.3.5
Unknown
The tet (U) gene produces a small
protein (a hundred and five amino acids) that confers low-stage tetracycline
resistance (Chopra and Roberts 2001). The TetU protein has 21% similarity over
its period to the TetM protein, but it does no longer include the consensus
GTP-binding sequences, which might be thought to be very critical for
tetracycline resistance in ribosomal safety proteins. The Tet (U) gene has been
identified in a vancomycin- and tetracycline-resistant S. aureus pressure that
did now not deliver the tet (k), Tet (L), tet (M), or tet (O) genes. From the
equal patient, vancomycin-resistant Enterococci were cultured that
carried both the tet (U) and tet (L) genes and some isolates also carried the
tet (ok) and/or Tet (M) genes (Weigel et al. 2004). The tet (U) gene has also
been diagnosed in Enterococcus spp. The importance of the Tet(U) gene is
uncertain since each Enterococcus and Staphylococcus isolates are
able to carry a style of efflux and ribosomal protection tet genes.
2.4
Sulfonamide Resistance Genes
The sulfonamides, the primary
antimicrobials developed for huge-scale creation into clinical practice (in
1935), goal dihydropteroate synthase. Their serendipitous discovery (the
antibacterial activity turned into visible initially in vivo while the active
compound changed into released as a part of a dye) pales handiest in contrast
with that of Fleming’s danger discovery of penicillin (Levy 2002). Two sul
genes (sulI and sulII) and one genetic element associated with mobile
antibiotic resistance genes [class 1 integron (intI1)] in 8 farm animals farms
in Hangzhou, jap China changed into investigated (Cheng et al. 2013).
2. 5
resistance charges and characteristics
Antibiotic opposition patterns of
integrity pathogens to the medication used to handle ruling class change
considerably with and within worldwide districts. Those dissimilarities are
compelled with the aid of marvelous patterns of medicine use, obvious concerning
a country with a disorder burden, differences in getting an effort to first-
and 2nd-line remedies, and a load of co-contaminations, particularly ularly
sickness, the human immunodeficiency bug (HIV), and infection (O’Neill 2014).
Resistance costs have additionally existed equated accompanying migratory
medicine use: within the United States of America, pierces of opposing E.
coli compared considerably accompanying migratory extreme happiness in
aminopenicillin and fluoroquinolone prescriptions, backward by way of 1 month (Cosmic
and others. 2012). any medicine-opposing infections, in addition to H.
influenzae in kids beneath five, have better humanness quotes distinguished
accompanying inclined contaminations (27 versus 7% humanness). but, this
manifold hazard of fate isn't forever prevalent: inside the case of
healthcare-befriended contaminations, medicine opposition does no longer
considerably increase death or distance of healing organization live because of
bloodstream contaminations or pneumonia (Lambert and others. 2011).
Antibiotic-opposing contaminations furthermore gifts to the fiscal burden on
healthcare buildings. In Europe, they profit and conceive.
1.
five billion euros annually, which involves healthcare costs and output
deficits (i.e., each direct and roundabout expense) (EMA and ECDC 2009). Inside
the United States, the occurring cost to the healthcare automobile is as much
as $20 billion, and fertility misfortunes total sporadic $35 billion (CDC
2013). Overdone-profits fields and international sites. inside the United
States of America, CDC (2013) has wanted that more 2 million contaminations and
23,000 passing are on account of medicine resistance every 12 months. In
Europe, an expected 25,000 end of life are happening from medicine-opposing
infections (EMA and ECDC 2009). Resistance of Streptococcus pneumoniae
invasive isolates to medicines has descended in the United States of America of
the western hemisphere, from 34 to 17% from 1999 to 2013 for penicillins, and
from 15 to eight% from 1999 to 2012 for second-generation cephalosporins. From
1999 to 2012, fighting against microclines extended from 23 to 34%,
nevertheless fluoroquinolone opposition waited solid, at 2%. Among E. coli
and k. pneumoniae isolates, opposition to 0.33-era cephalosporins and
fluoroquinolones inflated progressively: for 0.33-cycle cephalosporin fighting
in E. coli, from 2 to twelfth%, and in ok. pneumoniae, from 8 to 19%;
for fluoroquinolone opposition in E. coli, from five to 30%, and in k.
pneumoniae, from 7 to 18%. With E. faecium invasive isolates, vancomycin
fighting increased from 65 to 76%. as distinguished accompanying added
excessive-pay nations, the United States of America has better costs of
fighting many Gram-active microorganisms, amounting to VRE and MRSA (CDDEP
2015). Low- and middle-gain fields and worldwide parts k. pneumoniae is the
maximum usually submitted Gram-distressing bacterium in Asia and Africa,
constituting nearly half of all Gram-horrible contaminations in neonates. In
Asia, the middle opposition of. Pneumoniae to medicine become 94%, and to
cephalosporins, 84%; in Africa, it curves into 100 and 50%, individually.
Multidrug fighting came in 30% of traces in Asia and 75% of lines in Africa (Le
Doare and others. 2014). In sub-Saharan Africa, prices of multidrug opposition
surpassing 50% were noticed in obtrusive typhoidal and Nontyphoidal Salmonella
contaminations. Resistance to the drugs used to treat multidrug opposing Salmonella,
to a degree fluoroquinolones, is again growing (Kariuki and others. 2015).
Invasive nontyphoidal Salmonella contaminations are the reason for more
than 600,000 passing occurring, 55% of the ruling class in Africa (Kariuki and
others. 2015). Patterns of medicine opposition clash kind of in Latin America
and the Caribbean, place predominance of society-mixed Enterobacteriaceae
contaminations is above in the rest of the realm, especially in urinary area
contaminations induced by E. coli and event-intestinal contaminations
precipitated by E. coli and Klebsiella spp. These contaminations show
growing opposition to trimethoprim-sulfamethoxazole, quinolones, and second-creation
cephalosporins. In 2009, rates of opposition in urinary lot E. coli
isolates attained 71% in women and 85% in sons, accompanying the maximal rates happening
in Argentina and Peru (Salles and others. 2013). In Latin America and the Caribbean
in 2013, opposition in society S. pneumoniae isolates was mainly reduced to
penicillins but categorized from 0% in Bolivia to 97% in Chile. No fighting was
discovered to vancomycin, and very reduced opposition was discovered in some
nations after second-production cephalosporins. Resistance in E. faecium
nursing home isolates was above for E. faecalis. Resistance in E. faecium was
extreme to ampicillin and vancomycin, arriving 100% opposition to ampicillin in
Ecuador, El Salvador, and Paraguay. Paraguay likewise had the maximal fighting
to vancomycin, at 75%. E. faecalis fighting to medicine was categorized from 0
to 15%, and resistance to vancomycin was categorized from 0 to 22% (PAHO expected).
In Nepal, fighting rates surpassed 50% for S. pneumoniae and K.
pneumoniae isolates to usually secondhand situations, bearing raised from
2000 to 2008. Resistance of Salmonella typhi and Salmonella paratyphi
strains have still raised since 1998, and in E. coli, from 2006 to 2010.
Resistance rates were above 50% for all drugs proven in E. coli urinary
lot infections and extreme fighting rates were discovered in gonorrheal
Contaminations.
2.6
Global Patterns and Emerging Threats
The most current general estimates
of worldwide medicine opposition, written by the World Health Organization
(WHO) in 2014, list Escherichia coli, Klebsiella pneumoniae, and Staphylococcus
aureus as the three powers of excellent concern, guide two together
clinic- and community-captured contaminations. In five of the six WHO domains,
few nations stated E. coli opposition in addition to 50% to fluoroquinolones
and triennial-creation cephalosporins. K. pneumoniae opposition rates to
after second-production cephalosporins are above 30% private WHO appendage countries
and surpass 60% in a few domains (WHO 2014). MRSA opposition rates surpass 20%
comprehensively WHO domains and are above 80% in a few domains (WHO 2014).
Streptococcus pneumoniae,
nontyphoidal Salmonella, Shigella spp., and Neisseria gonorrhoeae
were more recognized as society-seized contaminations of high worldwide concern.
High rates of fighting first- and second-line drugs have previously increased
insult confidence in desperate remedy drugs, in the way that carbapenems (WHO
2014). This report supports a survey of the best accessible dossier on medicine
resistance rates general, illustration from Resistance Map (computer
network.resistancemap.org, a global computerized data in system of medicine use
and opposition news, developed for one Center fo Disease Dynamics, Economics
and Policy [CDDEP]), WHO, civil beginnings, and scientific news
Research
Method
The research proposed to accept the
hereditary mechanisms behind medicine fighting and the disposal of opposition
genes across miscellaneous bacterial strains. To achieve this, the following
arrangement was working:
Sample Collection: Bacterial strains
were calm from various sources, containing nursing home atmospheres, society
hospitals, and natural water crowds to capture a general of medicine-fighting
genes.
DNA Extraction and Preparation:
High-quality genomic DNA was derived using a patterned origin code. The
innocence and aggregation of DNA were confirmed utilizing spectrophotometry and
coagulate electrophoresis.
Whole-Genome Sequencing (WGS): DNA
samples were committed to extreme-throughput next-production sequencing (NGS)
to identify the ghost of medicine-fighting genes. Sequencing was acted using
Illumina or Pacific Biosciences podiums for extreme veracity and inclusion.
Bioinformatics
Analysis:
Gene Identification: Raw series data
was treated, uncluttered, and joined utilizing program tools like FASTQC and
Trimmomatic. Bioinformatics forms, containing BLAST (Basic Local Alignment
Search Tool) and Resistance Gene Identifier (RGI), were used to print and label
medicine resistance genes established popular databases to a degree CARD
(Comprehensive Antibiotic Resistance Database).
Linkage and Co-incident Analysis: A
Reasoning was performed to study the relation between fighting genes and their
potential unions with plasmids, transposons, and integrons utilizing forms like
BLASTn and custom-built handwriting in R.
Phenotypic Confirmation: To
substantiate the presence of recognized opposition genes, platter spread and
minimum inhibitory aggregation (MIC) tests were conducted to determine
bacterial opposition to medicines.
Statistical Analysis: Data from
deoxyribonucleic acid labeling and phenotypic testing were assembled and
resolved utilizing mathematical programs (e.g., SPSS, R). Chi-square and
equivalence tests were used to decide the partnership between various fighting
genes.
Results
The study recognized and
classification various antibiotic opposition genes and established their
historical linkages and the phenotypic characteristics they awarded:
Efflux
Genes:
Tet (A) and tet(B) genes were
commonly raised, connected to β-lactamase genes like blaTEM, and added fighting
causes such as strA and strB for medicine opposition.
Mer
operon commonly guides these genes, displaying resistance to the major planet.
Ribosomal
Protection Genes:
Tet (M) was outstanding with the
strains, providing medicine resistance through ribosomal guardianship.
Co-incident accompanying erm(B) was evident, deliberating resistance to MLSB
(macrolide-lincosamide-streptogramin B).
Enzymatic
Resistance Genes:
Tet (X) was labeled, connected to
MLSB fighting through allure catalyst activity. Other genes, to a degree aph
A-3 (kanamycin fighting), were again noticed.
Transposons
and Integrons:
Tn21,
Tn916, and Tn917 were raised to transfer data from one computer system to
another delivering genes and donating to the spread of medicine fighting.
Gene
Linkages:
Resistance genes were shown to
cluster together in the genome. Tet (A) repeatedly guided SGI1 (Salmonella
genomic archipelago 1) and plasmids, signifying potential transferability.
Genes
like Tet(G) were linked to aadA2 and dfrA genes, suggesting multi-drug
opposition sketches.
Discussion
The judgments underline the
complicatedness of antibiotic opposition systems and the duty of traveling
historical elements in their distribution:
Horizontal
Gene Transfer (HGT):
The study manifested the function of
plasmids, transposons, and integrons as instruments for deoxyribonucleic acid
transfer, promoting the spread of opposition genes like tet(A), tet(B), and
tet(M) across the bacterial public.
The attendance of Tn21 and Tn916
expedited the transfer of resistance genes, emphasizing the duty of movable
historical fundamentals in antimicrobial resistance (AMR).
Multidrug
Resistance:
The co-incident of genes in the way
that tet(A), blaTEM, and strA/B in sure strains displays the potential for
multi-drug fighting, which confuses situation methods and makes necessary
alternative healing approaches.
Implications
for Public Health:
The prevalence of medicine
opposition genes in two together dispassionate and referring to practices or
policies that do not negatively affect the environment samples points to the
urgent need for listening and attack. Resistance deoxyribonucleic acid
reservoirs in incidental sources can be a part of a hatchery for antimicrobial
opposition, conceivably moving human health.
Mechanisms
of Resistance:
The dossier disclosed various
methods of opposition, including outflow pumps (tet(A), tet(B)), ribosomal
guardianship (tet(M)), and concerned with atom and molecule change inactivation
(tet(X)). This variety in fighting mechanisms portrays the metamorphic
changeability of microorganisms.
Challenges
and Future Directions:
Identifying novel genes and
understanding their interplays with popular fighting causes are detracting from
developing inclusive AMR methods. Enhanced following schemes and the incidence
of new antibiotics mean opposing pathogens are alive.
Conclusion
This study focal points the complex
character of antibiotic-fighting genes and their part in the all-encompassing
energy challenge formal by antimicrobial resistance. The judgments disclose
that fighting genes are not only extensive but more often connected to
accompanying movable ancestral elements, promoting their speedy spread.
Strategies to combat AMR concede possibility involve exact surveillance, mean
interferences, and the growth of new medicines. Public health procedures should
supply instructions ruling the incidental spread of opposition genes and
advancing the rational use of medicines to maintain their productiveness for
future eras.
Acknowledgment:
The talent having to do with this research
project would not have occurred likely outside the abundant support and help of
entirety and plans. We do not any more our real recognition to all those
individuals who risked a function in the progress concerning this project. I at
this moment recognize that: I have no business-related or additional individual
interests, honestly or corner ways, in few matters that may influence or bias my
honesty as a person who writes about factual events for a living having to do
with this book.
Conflicts
of Interest:
The authors
declare that they have no conflicts of interest.
Financial
Support and Protection:
No extrinsic
capital for a project was captured to assist in accompanying the development of
this Manuscript
References
Yang,
S.-C., Chang, W.-J., Chang, Y.-H., Tsai, Y.-S., Yang, T.-P., Juan, C.-W. and
Shiau, M.-Y. "predominance of medicine fighting and OXA carbapenemase
genes in multidrug-opposing isolates of Acinetobacter baumannii
priceless Taiwan." Eu chronicle of experimental Microbiology &
Infectious ailments 29 (2010): 601–604.
Yun-Jian,
H. and Dong-ke, C. "2007 Mohnarin record: Antibiotic fighting of
non-fermenting Gram-negative bacterium." about the Orient Newspaper of
Antibiotics 33, no. 10 (2008): 597–601 + S6.
Bennett,
J.W., and K.-T. Chung. "Alexander Fleming and the lie of
Antibiotics." Advances in achieved Microbiology quadragesimal nine (2001):
163–184.
Shore,
L. and A. Pruden. "Launch." In Hormones and formula drugs Generated
by met Animal Feeding Operations, refined by way of L. Shore and A. Pruden,
1–five. The substantial sphere: Springer, 2009.
Rodriguez-Mozaz,
S., Chamorro, S., Marti, E., Huerta, B., Gros, M., Sanchez-Melsio, A., Borrego,
C.M., Barceló, D., and Balcázar, J.L. "predominance of Antibiotic Genes
and medicine fighting in hospital connected with university and concerning
cities wastewater and their effect at the taking waterway 69 (2015): 234–242.
Tang,
X., Lou, C., Wang, S., Lu, Y., Liu, M., Hashmi, M.Z., Liang, X., Li, Z., Liao,
Y., and Qin, W.' results of complete manure requests on the occurrence of
medicines and medicine opposition genes (ARGs) in edible grain soils: evidence
from four field experiments in Southeast China. Soil Biology and Biochemistry
90 (2015): 179–187.
Zhu,
Y.-G., Johnson, T.A., Su, J.-Q., Qiao, M., Guo, G.-X., Stedtfeld, R.D.,
Hashsham, S.A., and Tiedje, J.M. "differing and plentiful Antibiotic
Resistance Genes in about the orient Pig Farms." court cases of the
concerning a country with an Academy of Sciences united states of America
individual 110(2013): 3435–3440.
Martinez,
J.L. "Environmental contamination accompanying medicines and cause of
medicine fighting." Environmental pollution 157 (2009): 2893–2902.
Datta,
N. and V.M. Hughes. "Plasmids of the alike Inc groups in Enterobacteria
former than and afterward healing use of medicines." Nature 306 (1983):
616–617.
Akasaki,
k., Karasawa, k., Watanabe, M., Yonehara, H., and Umezawa, H.
"Monazomycin, a new medicine created by way of Streptomyces."
pamphlet of Antibiotics 16 (1963): 127–131.
Atkinson,
B. A., Abu-Al-Jaibat, A., and LeBlanc, D. J. "Antibiotic fighting among Enterococci
unique from experimental examples from 1953 to 1954." Antimicrobial powers
and Chemotherapy quadragesimal individual (1997): 1598–1600.
Cousin,
S. L., Jr., Whittington, W. L., and Roberts, M. C. "acquired macrolide
fighting genes and 1 bp deletions in the mtrR supporter in Neisseria
gonorrhoeae". Journal of Antimicrobial Chemotherapy having 50 of something
individual (2003): 131–133.
Roberts,
M. C. "change on acquired Antibiotic Resistance Genes." FEMS
Microbiology Letters 245 (2005): 195–203.
Pei,
R., Kim, S.-C., Carlson, ok. H., and Pruden, A. "impact of River Horizon
on Sediment Antibiotic Concentrations and Corresponding Antibiotic Resistance
Genes (ARGs)." Water studies 40 (2006): 2427–2435.
Kumar
Asamy, Toleman, M.A., Walsh, T.R., Bagaria, J., Butt, F., Balakrishnan, R.,
Chaudhary, U., Doumith, M., Giske, C.G. and Irfan, S. "Emergence of a new
device of medicine fighting in India, Pakistan, and the UK: a microscopic,
organic and epidemiological study." The Lancet Infectious Diseases 10
(2010): 597–602.
Levy,
S. "active Efflux: A dominant Mechanism for Biocide and Antibiotic
Resistance." periodical of completed activity Microbiology ninety-two
(2002): 65S–71S.
Roberts,
M. C. "Tetracycline fighting by way of ribosomal security proteins."
In Frontiers of Antimicrobial Resistance, 19.–28. Washington, DC: American
Society for Microbiology, 2005.
Thaker,
M., Spanogiannopoulos, P., and Wright, G.D. "Antibiotic opposition".
Container and Molecular History Sciences 67 (2010): 419–431.
Roberts,
M. C. "Mechanisms of Bacterial Antibiotic Resistance and communication
discovered from herbaceous Antibiotic-Resistant Microorganisms." In
Antimicrobial Resistance in the Environment, 93–121. Washington, DC: American
Society of Microbiology, 2011. https://doi.org/10.1002/9781118156247.ch7.
Levy,
S. B. "Mechanisms for opposition in soil." concerning details talent
312 (2006): 529.
![]() Antibiotic resistance has become one
of the most pressing worldwide fitness issues, jeopardizing the effectiveness
of contemporary medication. Resistance genes, commonly observed in cellular
genetic factors such as plasmids, transposons, and integrons, are imperative to
the spread of resistance across bacterial populations. Those genes allow the
microorganism to continue to exist with exposure to antibiotics, rendering
well-known treatments useless. The overuse and misuse of antibiotics in human
medication, agriculture, and veterinary practices have contributed
significantly to the fast emergence of resistant lines. The resistance
mechanisms encompass antibiotic degradation by enzymes, modification of
antibiotic objectives, reduced drug uptake, and activation of efflux pumps that
expel antibiotics from the bacterial cell. Horizontal gene transfer (HGT),
through approaches that include conjugation, transformation, and transduction,
permits the big distribution of resistance genes across bacterial species,
enhancing their patience in the environment. The continuous movement of
resistance genes among people, animals, and the environment complicates efforts
to govern resistance. Advances in molecular biology techniques, such as
subsequent-generation sequencing, have facilitated the identification and
characterization of resistance genes, offering precious insights into their
diversity, evolution, and capacity reservoirs. Environmental monitoring has revealed
the presence of resistance genes in various ecosystems, including water, soil,
flora, and fauna, emphasizing the interconnected nature of human, animal, and
environmental health referred to as the only health technique. Combating
antibiotic resistance requires a coordinated, multidisciplinary effort that
integrates surveillance, stewardship, and the improvement of novel healing
techniques.
Antibiotic resistance has become one
of the most pressing worldwide fitness issues, jeopardizing the effectiveness
of contemporary medication. Resistance genes, commonly observed in cellular
genetic factors such as plasmids, transposons, and integrons, are imperative to
the spread of resistance across bacterial populations. Those genes allow the
microorganism to continue to exist with exposure to antibiotics, rendering
well-known treatments useless. The overuse and misuse of antibiotics in human
medication, agriculture, and veterinary practices have contributed
significantly to the fast emergence of resistant lines. The resistance
mechanisms encompass antibiotic degradation by enzymes, modification of
antibiotic objectives, reduced drug uptake, and activation of efflux pumps that
expel antibiotics from the bacterial cell. Horizontal gene transfer (HGT),
through approaches that include conjugation, transformation, and transduction,
permits the big distribution of resistance genes across bacterial species,
enhancing their patience in the environment. The continuous movement of
resistance genes among people, animals, and the environment complicates efforts
to govern resistance. Advances in molecular biology techniques, such as
subsequent-generation sequencing, have facilitated the identification and
characterization of resistance genes, offering precious insights into their
diversity, evolution, and capacity reservoirs. Environmental monitoring has revealed
the presence of resistance genes in various ecosystems, including water, soil,
flora, and fauna, emphasizing the interconnected nature of human, animal, and
environmental health referred to as the only health technique. Combating
antibiotic resistance requires a coordinated, multidisciplinary effort that
integrates surveillance, stewardship, and the improvement of novel healing
techniques.