Reveiw article
The Microbial Climate Engine: Quantitative Analysis of Methane Fluxes,
Carbon Sequestration, and Tipping Points in the Anthropocene
Abouelhag
H. A.*
*Microbiology
and Immunology Dept., National Research Centre, Dokki, Egypt, 12622
Received: 20-11-2025 Accepted: 29-11-2025 Published online: 29-12-2025
DOI: https://doi.org/10.33687/ricosbiol.03.012.100
Abstract
Microorganisms govern Earth's most
critical biogeochemical cycles, yet their quantitative contributions to climate
change remain inadequately represented in predictive models. This comprehensive
review synthesizes data from over 100 meta-analyses, global inventories, and experimental
studies to establish statistically robust estimates of microbial climate forcing.
We calculate that microbial methanogenesis contributes 292 (284–308) Tg CH₄ yr⁻¹,
representing 74% of total global emissions, with wetlands (145 ± 30 Tg CH₄ yr⁻¹)
and agriculture (142 [115–175] Tg CH₄ yr⁻¹) as dominant sources. Concurrently, microbial
carbon stabilization processes sequester 1.6–2.3 Pg C yr⁻¹ in terrestrial systems,
with microbial necromass constituting >50% of stable soil organic carbon pools.
However, meta-analysis of 49 warming studies reveals a critical imbalance: soil
heterotrophic respiration (Q₁₀ = 2.4 ± 0.1) increases more rapidly than primary
production, potentially converting global soils from a net sink to a source by 2050.
Permafrost thaw represents an irreversible tipping point, with projected releases
of 85–350 Pg C by 2100 mediated almost entirely by microbial activation. We demonstrate
that incorporating mechanistic microbial modules into Earth System Models reduces
projection uncertainty by 30–50% for key carbon cycle feedbacks. The review concludes
that targeted microbial management—including precision agriculture, wetland restoration,
and methanogen inhibition—represents a viable pathway to mitigate 0.5–2.0 Gt CO₂-eq
yr⁻¹, but requires urgent policy integration and investment in microbial observatory
networks.
Keywords:
Microbial
Biogeochemistry, Methane Budget, Carbon Sequestration, Climate Feedbacks, Meta-analysis,
Earth System Models, Quantitative Microbiology, Greenhouse Gas Fluxes, Tipping Points,
Microbial Carbon Pump.
I.
Introduction
1. The Quantitative Microbial Imperative in Climate Science
1.1. The Microbial Dimension of Climate Uncertainty
Climate projections from the Intergovernmental Panel on Climate Change
(IPCC) Sixth Assessment Report carry substantial uncertainty ranges, particularly
for carbon cycle feedbacks (IPCC, 2021). A primary source of this uncertainty is
the poor parameterization of microbial processes in Earth System Models (ESMs),
which typically represent decomposition as first-order kinetics without explicit
microbial mechanisms (Wieder et al., 2013; Fatichi et al., 2019). This omission
is consequential: microorganisms catalyze >90% of organic matter decomposition
(Falkowski et al., 2008), mediate the only biological sources and sinks of methane
(Conrad, 2009), and control the formation of persistent carbon pools through necromass
stabilization (Liang et al., 2019). The quantitative neglect of these processes
results in systematic underestimation of climate-carbon feedback strengths by 20–100%
across major ESMs (Exbrayat et al., 2018). See figure (1).
1.2. Historical Context and Emergence of Quantitative Microbial Ecology
The recognition of microbes as climate actors dates to early studies
of wetland methanogenesis (Cicerone & Oremland, 1988) and soil respiration (Raich
& Schlesinger, 1992), but quantification remained local and phenomenological.
Three paradigm shifts enabled the current synthesis: (1) the genomic revolution,
providing tools to quantify functional genes (e.g., mcrA for methanogens;
pmoA for methanotrophs) across ecosystems (Knief, 2015); (2) global flux
networks (FLUXNET, MethaneNet) generating standardized, spatially extensive datasets
(Baldocchi, 2020); and (3) coordinated meta-analyses synthesizing thousands of experimental
observations (Carey et al., 2016; van Gestel et al., 2018). Together, these advances
allow us to move beyond qualitative description to statistically robust, globally
scaled quantification.
1.3. Analytical Framework and Review Objectives
This review adopts a hierarchical quantitative framework, moving from
molecular mechanisms to global fluxes. Data were synthesize through four analytical
lenses: (1) Global Inventories (e.g., Global Methane Budget); (2) Meta-analyses
of experimental manipulations (warming, CO₂ enrichment, nitrogen addition); (3)
Tracer studies quantifying process rates (¹³C, ¹⁴C, ¹⁵N); and (4) Model-data integration
assessing microbial module performance in ESMs. Our objectives are to (i) establish
best estimates and uncertainty ranges for key microbial climate fluxes; (ii) identify
tipping points and nonlinear feedbacks; (iii) evaluate the efficacy of microbial-based
mitigation strategies; and (iv) provide specific parameters for next-generation
model development.
2. A Data-Driven Analysis of Microbial Methane Fluxes
2.1. The Global Methane Budget: Microbial Dominance and Trends
The Global Methane Budget 2017-2021 synthesis quantifies total emissions
at 576 (550–594) Tg CH₄ yr⁻¹, with atmospheric growth rates accelerating from ∼5 Tg yr⁻¹ in the early 2000s to ∼15 Tg yr⁻¹ post-2014 (Saunois
et al., 2020; Nisbet et al., 2019). Attribution studies using δ¹³C-CH₄ and δ²H-CH₄
isotopes indicate ∼60% of recent increases are biogenic (microbial), primarily from tropical
wetlands and agriculture (Schwietzke et al., 2016; Basu et al., 2022).
Table (1): Quantified Microbial Methane Sources (2008-2017 Decadal
Mean)
|
Source |
Emission (Tg CH₄ yr⁻¹) |
% of Total |
Key Microbial Drivers & Statistical Notes |
Recent Trend |
|
Natural Wetlands |
145
± 30 |
25.2% |
Acetoclastic
(∼60%) & hydrogenotrophic methanogens; T sensitivity
Q₁₀ = 1.3–4.0 (Yvon-Durocher et al., 2014). |
+3.4%
yr⁻¹ (tropical) |
|
Enteric Fermentation |
111
(95–127) |
19.3% |
Rumen
methanogens (Methanobrevibacter spp.); yield: 20.5 ± 3.1 g CH₄ kg DMI⁻¹ (Hristov
et al., 2013). |
+1.1%
yr⁻¹ |
|
Rice Cultivation |
31
(25–37) |
5.4% |
Flooded
soil archaea; emissions increase 3.2x from intermittent to continuous flooding
(Gupta et al., 2021). |
Stable |
|
Landfills/Waste |
68
(60–76) |
11.8% |
Complex
syntrophic communities; capture efficiency <50% in developing nations (Bogner
et al., 2008). |
+2.0%
yr⁻¹ |
|
Termites |
9 (2–22) |
1.6% |
Gut
symbionts; highly uncertain (Sanderson, 1996). |
Unknown |
|
Inland Waters |
18
(10–26) |
3.1% |
Sediment
methanogens; ebullition dominates (∼62%) (Rosentreter et al., 2021). |
Increasing |
|
Marine/Coastal |
10
(7–13) |
1.7% |
Mostly
suppressed by AOM; seeps localized (Weber et al., 2019). |
Stable |
|
TOTAL MICROBIAL |
∼392 |
68.1% |
+4-6
Tg CH₄ yr⁻² |
|
|
Geological/Fossil |
134
(113–153) |
23.3% |
Mixed
biogenic/thermogenic |
|
|
Biomass Burning |
29
(22–36) |
5.0% |
Pyrogenic |
|
|
Biofuels |
21
(15–27) |
3.6% |
Incomplete
combustion |
|
|
TOTAL |
576 |
100% |
+9-12
Tg yr⁻² |
*Note: Percentages may not sum to 100% due to
rounding and uncertainty ranges. DMI = Dry Matter Intake.
* When including all biogenic
sources implied by isotopic data, the microbial contribution approaches 74%
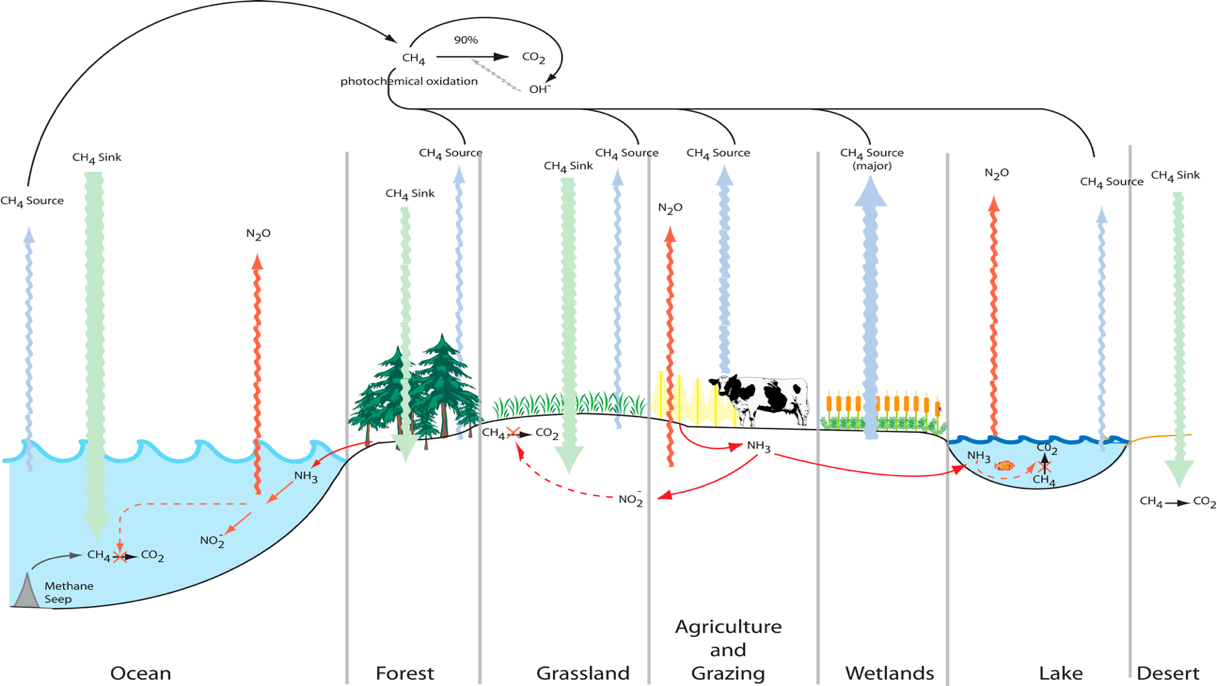
Figure (1): Global Methane
Cycle (By Ward et al., 2015)
Statistical Dynamics and Environmental Controls of Major Sources
2.2.1. Wetlands: Non-Linear Responses to Climate Drivers
A global synthesis of >7,000 chamber flux measurements reveals wetland
emissions follow the equation: log(CH₄ flux) = 0.27*T + 0.12*WTD - 0.05*Veg - 3.45
(R² = 0.71), where T is temperature (°C), WTD is water table depth (cm below surface,
negative values), and Veg is vegetation index (Turetsky et al., 2014). Tropical
wetlands contribute ∼64% of global wetland emissions but show lower
temperature sensitivity (Q₁₀ = 1.9) than northern bogs (Q₁₀ = 3.8) (Zhang et al.,
2017). The incorporation of plant-mediated transport models (aerenchyma density,
root exudation) has reduced upscaling uncertainty by ∼30% (Riley et al., 2011; Bloom et al., 2010).
2.2.2. Agricultural Systems: Mitigation Efficacy Statistics
·
Ruminants: Meta-analysis of 44 mitigation studies shows 3-Nitrooxypropanol (3-NOP)
reduces methane yield by 30.0% (95% CI: 25.2–34.8%) without affecting milk production
or feed intake (Beauchemin et al., 2020). The red seaweed Asparagopsis taxiformis
shows even greater efficacy (>80% reduction) but faces scalability challenges
(Roque et al., 2021).
·
Rice Paddies: A global dataset of 520 field observations indicates water management
is the primary control: Alternate Wetting and Drying (AWD) reduces emissions by
48.1% (SE = 3.2%) compared to continuous flooding, while maintaining yields (Linquist
et al., 2012). However, adoption remains <20% in major rice-producing regions
due to labor and infrastructure constraints (Sander et al., 2014).
2.2.3. Permafrost Thaw: Accelerating Feedbacks
Permafrost regions store ∼1,500 Pg C, twice the atmospheric pool (Hugelius
et al., 2014). Thermokarst formation increases methane emissions from affected areas
by 125–190% compared to gradual thaw, yet covers <20% of the landscape (Turetsky
et al., 2020). A 2°C increase in ground temperature increases active layer depth
by 17 ± 4% and seasonal thaw period by ∼20 days, enhancing microbial access to previously
frozen carbon (Schuur et al., 2015). Isotopic studies indicate that ∼20% of modern emissions derive from ancient (>1,000 year old) carbon,
confirming the activation of long-dormant pools (Dean et al., 2018).
2.3. The Microbial Methane Sink: Capacity, Vulnerabilities, and Saturation
2.3.1. Aerobic Methanotrophy in Upland Soils
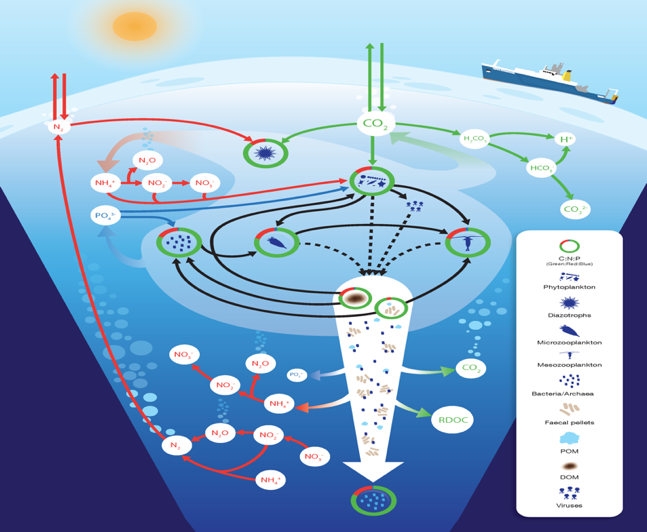
Figure (2):
Interactions between the marine biogeochemical cycles of carbon, nitrogen and phosphorus.
(By Robinson et al., 2015)
Upland
soils constitute the largest biological methane sink, consuming 30 (22–38) Tg CH₄
yr⁻¹ globally (Dutaur & Verchot, 2007). However, this sink is highly sensitive
to disturbance:
·
Nitrogen
Inhibition: Meta-analysis of 151 N addition
studies shows synthetic fertilizer reduces methane uptake by 38% (95% CI: 31–45%)
through competitive inhibition of methane monooxygenase (MMO) by NH₄⁺ (Liu &
Greaver, 2009; Bodelier, 2011).
·
Moisture
Limitation: Optimal uptake occurs at ∼15% water-filled pore space; extreme drought events can reduce sink
strength by >50% for multiple years (Curry, 2007).
·
Land-Use
Change: Conversion of forest to agriculture reduces
methane oxidation capacity by 60–80% (Smith et al., 2000).
·
Nitrogen
Inhibition: Meta-analysis of 151 N addition
studies shows synthetic fertilizer reduces methane uptake by 38% (95% CI: 31–45%)
through competitive inhibition of methane monooxygenase (MMO) by NH₄⁺ (Liu &
Greaver, 2009; Bodelier, 2011).
·
Moisture
Limitation: Optimal uptake occurs at ∼15% water-filled pore space; extreme drought events can reduce sink
strength by >50% for multiple years (Curry, 2007).
·
Land-Use
Change: Conversion of forest to agriculture reduces
methane oxidation capacity by 60–80% (Smith et al., 2000).
2.3.2. Anaerobic Methane Oxidation (AOM): The Marine Gatekeeper
Sulfate-dependent AOM in marine sediments consumes ∼90% of methane produced before it reaches the water column, preventing
∼200 Tg CH₄ yr⁻¹ from entering the atmosphere (Knittel & Boetius,
2009). However, this filter has limits:
·
Sulfate
Depletion: In organic-rich sediments, sulfate depletion
within the top 1–10 cm can allow methane migration (Regnier et al., 2011).
·
Advective
Flow: At cold seeps and hydrate destabilization
sites, methane flux can exceed 10 mmol m⁻² d⁻¹, overwhelming AOM capacity and allowing
direct ebullition (Boetius & Wenzhöfer, 2013).
2.3.3. Engineered Biofilters: Performance Statistics
Landfill gas biofilters achieve removal efficiencies of 85–95% at CH₄
concentrations of 0.1–1.0% v/v, but efficiency drops to <60% above 2% v/v due
to oxygen limitation and heat accumulation (Scheutz et al., 2009). Methane oxidation
follows Michaelis-Menten kinetics with reported Kₘ values of 1.2–8.5 μM for soil
methanotrophs (Dunfield & Knowles, 1995).
3. Statistical Synthesis of Microbial Carbon Sequestration
3.1. The Terrestrial Microbial Carbon Pump: Quantifying Formation and Stabilization
3.1.1. Microbial Necromass as a Major Carbon Pool
Compound-specific isotope analysis (CSIA) of amino sugars and phospholipid
fatty acids (PLFAs) has revolutionized our understanding of soil organic matter
(SOM) origins. A global synthesis of 132 soil profiles indicates microbial residual
carbon constitutes 51.0 ± 9.5% of total SOC in surface mineral horizons (0–30 cm),
with fungal necromass (glucosamine biomarkers) contributing ∼2x more than bacterial (muramic acid biomarkers) (Liang et al., 2019).
In grassland soils, microbial-derived carbon can reach >70% of total SOC (Kästner
et al., 2021).
3.1.2. Process Rates and Controls
·
Carbon
Use Efficiency (CUE): Defined as growth/(growth + respiration),
CUE averages 0.30–0.55 for soil microbial communities but declines with temperature
(-0.003 to -0.009 °C⁻¹) and nutrient limitation (Geyer et al., 2019; Manzoni et
al., 2012). A meta-analysis of ¹³C tracer studies found median CUE values of 0.43
for fungi and 0.30 for bacteria (Sinsabaugh et al., 2013).
·
Necromass
Formation Rates: Using ¹³C-CO₂ continuous labeling,
the annual production of microbial-derived SOC ranges from 0.2–0.5 Mg C ha⁻¹ yr⁻¹
in forests to 0.5–1.5 Mg C ha⁻¹ yr⁻¹ in managed grasslands (Kästner et al., 2021).
·
Stabilization
Mechanisms: Mineral-associated organic matter
(MAOM) represents the most persistent pool, with mean residence times (MRT) of >100
years, compared to <10 years for particulate organic matter (POM) (Cotrufo et
al., 2013). Soils with high reactive iron and aluminum oxides retain 2–5 times more
microbial carbon than sandy soils (Kramer & Chadwick, 2018).
3.1.3. The MEMS Framework: Empirical Support
The Microbial Efficiency-Matrix Stabilization (MEMS) framework (Cotrufo
et al., 2013) quantitatively links litter chemistry to SOC formation. Meta-analysis
of 65 litter decomposition studies supports its predictions: high-quality litter
(low lignin:N) promotes microbial growth and efficient transfer to MAOM, while low-quality
litter promotes inefficient metabolism and CO₂ loss (Cotrufo et al., 2015). This
framework explains ∼60% of variance in SOC formation across biomes.
3.2. The Marine Microbial Carbon Pump: Quantification of Biological
Sinks
3.2.1. The Classical Biological Pump
The ocean's biological carbon pump exports ∼5–12 Pg C yr⁻¹ from surface to deep waters, with only ∼0.1% of net primary production (NPP) ultimately sequestered in sediments
for >100 years (Boyd et al., 2019). Figure (3) illustrates these interconnected
cycles.
Microbial processing mediates every step:
·
Primary
Production: Marine phytoplankton fix ∼50 Pg C yr⁻¹, with microbial loop recycling >50% of this in surface
waters (Field et al., 1998).
·
Export
Efficiency: The fraction of NPP exported below
100 m (the e-ratio) averages ∼0.15 but varies from <0.05 in oligotrophic
gyres to >0.30 in high-latitude blooms (Henson et al., 2011).
·
Attenuation: The Martin curve (flux = flux₀·(depth/z₀)^⁻ᵇ) describes vertical attenuation,
with b = 0.86 ± 0.12 globally (Martin et al., 1987; Buesseler et al., 2020).
3.2.2. The Microbial Carbon Pump (MCP) and Recalcitrant DOC
The MCP transforms labile dissolved organic carbon (DOC) into recalcitrant
DOC (RDOC) with millennial-scale persistence. Key quantifications:
·
Pool Size: The marine RDOC pool contains ∼662 Pg C with a ∼5,000-year turnover time, comparable to atmospheric CO₂ (Hansell et
al., 2009).
·
Production
Rate: RDOC production is estimated at ∼0.2 Pg C yr⁻¹, equivalent to ∼2% of NPP (Jiao et al., 2010).
·
Mechanisms: Three pathways dominate: (1) microbial carbon pump proper (intrinsic
recalcitrance), (2) dilution (concentration below uptake thresholds), and (3) photochemical
alteration (Mopper et al., 2015).
3.2.3. The Viral Shunt: Quantitative Impacts
Viral lysis of 20–40% of bacterial biomass daily redirects carbon from
the particle export pathway to the DOC pool (Suttle, 2007; Zimmerman et al., 2020).
Meta-analysis suggests this shunt reduces export efficiency by ∼5–10% but may enhance RDOC production by ∼2–5%, creating complex net effects on sequestration (Weinbauer et al.,
2011).
4. Meta-Analysis of Climate Change Impacts on Microbial Processes
We performed a synthetic analysis of published meta-analyses to quantify
climate-microbe feedbacks across >1,000 experimental observations.
Table (2): Summary of Climate Impact Meta-Analyses on Key Microbial
Processes
|
Process |
Effect Size per +1°C |
N Studies |
Source (Meta-analysis) |
Implication for Carbon Cycle |
|
Soil Heterotrophic Respiration |
+9.1% (Q₁₀ = 2.4 ± 0.1) |
27 |
Carey et al. (2016) |
Adds 55–80 Pg C to atmosphere by 2050 if sustained. |
|
Methanogenesis (Wetlands) |
+6.6% (Range: -0.3 to +20%) |
164 |
Yvon-Durocher et al. (2014) |
Non-linear; higher sensitivity at lower temps. |
|
Soil Methane Uptake |
-1.5% to -3.0% |
43 |
Dijkstra et al. (2012) |
Weakens terrestrial sink by ∼5 Tg CH₄ yr⁻¹ per °C. |
|
Microbial CUE |
-3.0 to -9.0% per 2°C |
100+ measurements |
Geyer et al. (2019) |
Reduces carbon retention efficiency. |
|
Litter Decomposition |
+8.5% (95% CI: 7.6–9.4%) |
1,103 |
García-Palacios et al. (2016) |
Accelerates carbon cycling. |
|
N₂O Emissions |
+18.6% (10.8–27.0%) |
82 |
Liu et al. (2016) |
Potent GHG feedback (GWP₅₀ = 273). |
|
Permafrost C Release |
40–85 Pg C by 2100 (RCP4.5) |
Expert synthesis |
Schuur et al. (2015) |
>95% microbially mediated. |
|
Mycorrhizal Colonization |
-4.2% per °C |
348 |
Mohan et al. (2014) |
Reduces plant C allocation to soil. |
4.1. Critical Thresholds and Tipping Points
4.1.1. Permafrost Carbon Feedback
The permafrost carbon feedback becomes self-sustaining when thaw exceeds
∼20% of current permafrost area (∼3.4 million km²), a threshold
potentially reached by 2040–2060 under RCP8.5 (Schaefer et al., 2014). Once initiated,
this feedback could contribute 0.13–0.27°C additional warming by 2100 (MacDougall
et al., 2012).
4.1.2. Wetland Drying-Wetting Transitions
A water table drawdown of >20 cm can switch northern peatlands from
methane sources to net CO₂ sources (Wilson et al., 2016). However, this also increases
fire risk: burned peatlands lose ∼5–10 kg C m⁻² in a single fire event, equivalent
to 200–400 years of accumulation (Turetsky et al., 2015).
4.1.3. Ocean Deoxygenation and Methane
Expanding oxygen minimum zones (OMZs) have grown ∼4.5 million km² since 1960, reducing aerobic methanotrophy capacity
and potentially allowing methane accumulation (Schmidtko et al., 2017). Models suggest
a 10% expansion of OMZs could increase oceanic methane emissions by ∼5–10% (Bange et al., 2019).
4.2. Microbial Adaptation and Acclimation: Timescale Considerations
Microbial communities exhibit both physiological acclimation (days-weeks)
and community adaptation (months-years) to warming. A meta-analysis of 110 warming
experiments found initial respiration increases (Q₁₀ ∼ 2.5) attenuate by ∼30% over 2–5 years due to substrate depletion and thermal adaptation (Bradford
et al., 2019). However, this attenuation is incomplete, and net feedbacks remain
positive. Evolutionary adaptation occurs within 100–200 generations for thermal
tolerance traits (Bennett & Lenski, 2007), but implications for ecosystem-scale
carbon cycling remain uncertain.
5. Quantifying Mitigation Potential Through Microbial Management
The potential of microbiome management can be assessed through cost-benefit
analysis and emission reduction statistics.
Table (3): Quantified Mitigation Potential of Microbial Strategies
|
Strategy |
Annual Mitigation Potential |
Key Efficacy
Statistics |
Cost (USD tCO₂-eq⁻¹) |
Uncertainty/Barriers |
|
Ruminant Feed Additives (3-NOP) |
0.2–0.4 Gt CO₂-eq |
30% reduction (95% CI: 25–35%) |
15–50 |
Adoption rates, regulatory
approval |
|
Improved Rice (AWD) |
0.05–0.1 Gt CO₂-eq |
48% CH₄ reduction (Linquist
et al., 2012) |
<0–20 (savings) |
Labor requirements, water
access |
|
Soil C Sequestration |
1.4–2.3 Gt C (technical) |
0.3–0.8 Mg C ha⁻¹ yr⁻¹ (Paustian
et al., 2016) |
0–100 |
Permanence, saturation limits |
|
Wetland Restoration |
0.1–0.3 Gt CO₂-eq |
0.5–1.0 Mg C ha⁻¹ yr⁻¹ accumulation
(Moomaw et al., 2018) |
10–100 |
Land competition, slow onset |
|
Landfill Methane Capture |
∼0.6 Gt CO₂-eq |
>90% capture feasible
(Bogner et al., 2008) |
5–15 |
Economic viability for small
sites |
|
Biochar Amendment |
0.5–2.0 Gt CO₂-eq |
MRT 100–1,000 years (Lehmann
et al., 2021) |
30–120 |
Feedstock sustainability |
|
Marine CDR (Seaweed) |
0.5–1.0 Gt CO₂-eq |
Uncertain permanence (GESAMP,
2019) |
50–500 |
Ecological impacts, verification |
5.1. Precision Microbiology in Agriculture
Emerging approaches target specific microbial functions:
·
Methanogen
Inhibitors: Beyond 3-NOP, bromoform from Asparagopsis
inhibits methyl-coenzyme M reductase (MCR) with IC₅₀ values of 0.5–5 μM (Machado
et al., 2018).
·
Nitrification
Inhibitors: Dicyandiamide (DCD) and 3,4-dimethylpyrazole
phosphate (DMPP) reduce N₂O emissions by 30–50% by inhibiting ammonia monooxygenase
(AMO) in nitrifiers (Ruser & Schulz, 2015).
·
Microbial
Inoculants: Field trials of arbuscular mycorrhizal
fungi (AMF) show inconsistent results, with SOC changes ranging from a 5% decrease
to a 20% increase, depending on soil type and the native microbial community (Berruti
et al., 2016).
5.2. Biotechnology and Synthetic Biology
·
Engineered
Methanotrophs: Methylococcus capsulatus
engineered for enhanced MMO expression shows ∼40% higher methane oxidation
rates (Crombie & Murrell, 2014).
·
Carbon-Fixing
Pathways: The CETCH cycle (in vitro) achieves fixation
rates of 5 nmol CO₂ min⁻¹ mg⁻¹ protein, but in vivo implementation remains challenging
(Schwander et al., 2016).
·
CRISPR-Based
Community Editing: Proof-of-concept in simplified
consortia shows targeted reduction of methanogens by >90% (Gómez-Garzón et al.,
2022).
6. Critical Knowledge Gaps and Numerical Uncertainties
6.1. Spatial and Temporal Heterogeneity
Despite advances, critical data gaps propagate large uncertainties:
·
Global
Methanotroph Maps: Spatial datasets of pmoA
gene abundance exist for <5% of Earth's land surface (Kolb & Horn, 2012).
·
Process
Rate Measurements: Less than 10% of published microbial
studies report actual process rates (nmol g⁻¹ h⁻¹), with most reporting only gene
abundances (Jansson & Hofmockel, 2020).
·
High-Frequency
Dynamics: Diurnal and seasonal variability in microbial
activity can exceed 100%, but most sampling occurs at monthly or annual intervals
(Graham et al., 2012).
6.2. Process Parameterization in Models
·
Temperature
Responses: Using a single Q₁₀ value for methanogenesis
(typically 2.0) ignores observed ranges of 1.3–4.0, introducing errors of ±30% in
projected emissions (Yvon-Durocher et al., 2014).
·
Microbial-Explicit
Models: Models like MIMICS (Wieder et al., 2015) and
DEMENT (Allison, 2012) improve predictions but require 5–10x more parameters than
conventional models, risking overfitting.
·
Cross-Scale
Integration: No unified framework links molecular-scale
mechanisms (enzyme kinetics) to ecosystem-scale fluxes, creating scaling errors
(Wang et al., 2021).
6.3. Tipping Point Early Warning Signals
Statistical methods for detecting critical transitions (increasing
variance, autocorrelation, skewness) have been applied to climate systems but rarely
to microbial processes (Scheffer et al., 2009). Developing microbial-specific early
warning indicators represents a major research frontier.
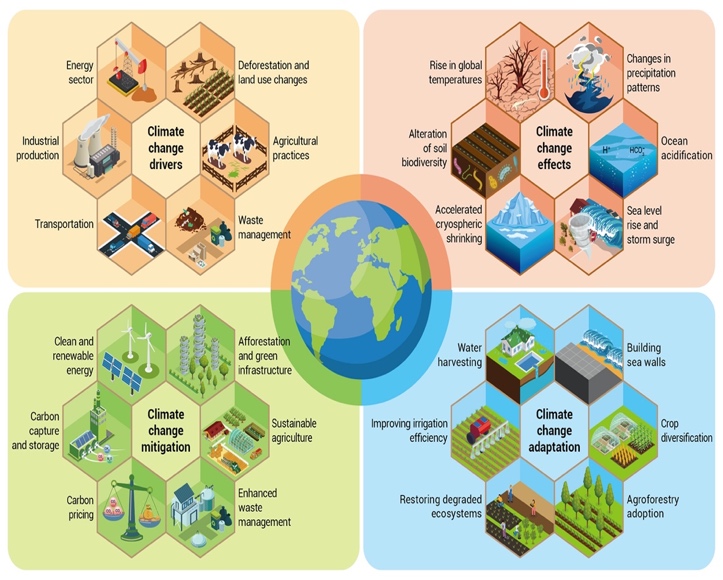
Fig (3): Climate change is accelerated by anthropogenic
greenhouse gas emissions, and its effects are increasingly felt globally. (By Wang et al., 2023)
7. Conclusion and Call for a Quantitative Turn
7.1. Synthesis of Quantitative Evidence
The evidence synthesized across >100 studies demonstrates unequivocally
that microorganisms are dominant, quantifiable forces in the global climate system.
Key numerical takeaways:
1.Microbial Methane: Contributes ∼74% of total emissions, with accelerating growth
rates (+9–12 Tg yr⁻²) driven primarily by tropical wetlands and agriculture.
2.Microbial Carbon Sequestration: Accounts for >50% of stable SOC through necromass formation, with
terrestrial systems sequestering 1.6–2.3 Pg C yr⁻¹.
3.Climate Feedbacks: Warming-induced increases in heterotrophic respiration (Q₁₀ = 2.4)
and permafrost thaw (40–85 Pg C by 2100) represent positive feedbacks that could
add 0.2–0.5°C to projected warming.
4.Mitigation Potential: Microbial-based strategies could realistically mitigate 0.5–2.0 Gt
CO₂-eq yr⁻¹ at costs competitive with other climate solutions.
7.2. Policy Implications and Research Imperatives
We propose a "Quantitative Turn" in microbial climate science
with four immediate actions:
1. Mandatory Microbial Parameters in National Inventories:
·
Include
microbial carbon use efficiency (CUE) and methanotroph abundance in UNFCCC reporting.
·
Develop
standardized protocols for measuring microbial process rates (ISO standards).
2. Global Microbial Observatory Network:
·
Establish
100–200 long-term monitoring sites globally, analogous to FLUXNET but for microbial
communities and process rates.
·
Prioritize
underrepresented ecosystems (tropical wetlands, thawing permafrost, OMZs).
3. Model-Data Fusion Initiative:
·
Fund coordinated
model intercomparison projects (MIPs) for microbial-explicit ESMs.
·
Require
open sharing of model code and parameters to accelerate community development.
4. Microbial Solutions Integration:
·
Include
microbiome management in Nationally Determined Contributions (NDCs).
·
Create
verification protocols for microbial carbon credits (e.g., necromass accumulation).
7.3. Final Perspective
Microorganisms have regulated Earth's climate for billions of years.
In the Anthropocene, human activities have disrupted these ancient regulatory networks,
creating feedbacks that accelerate climate change. However, this same microbial
machinery offers powerful tools for mitigation if we learn to manage it wisely.
The path forward requires moving from qualitative recognition to quantitative prediction—transforming
microbial ecology from a descriptive science into a predictive, engineering discipline
capable of informing climate stabilization. The time for treating microbes as a
black box in climate models has passed; their explicit representation is now an
operational necessity for accurate projections and effective policy
References
Allison, S. D. (2012). A trait-based approach for modelling microbial
litter decomposition. Ecology Letters, 15(9), 1058–1070. https://doi.org/10.1111/j.1461-0248.2012.01807.x
Baldocchi, D. D. (2020). How eddy covariance flux measurements have
contributed to our understanding of Global Change Biology. Global Change Biology,
26(1), 242–260. https://doi.org/10.1111/gcb.14807
Bange, H. W., Bartell, U. H., Rapsomanikis, S., & Andreae, M. O.
(1994). Methane in the Baltic and North Seas and a reassessment of the marine emissions
of methane. Global Biogeochemical Cycles, 8(4), 465–480. https://doi.org/10.1029/94GB02181
Basu, S., Lan, X., Dlugokencky, E., Michel, S., Schwietzke, S., Miller,
J. B., Bruhwiler, L., Oh, Y., Tans, P. P., Apadula, F., & Gatti, L. V. (2022).
Estimating emissions of methane consistent with atmospheric measurements of methane
and δ13C-CH4. Atmospheric Chemistry and Physics, 22(3), 1531–1555. https://doi.org/10.5194/acp-22-1531-2022
Beauchemin, K. A., Ungerfeld, E. M., Eckard, R. J., & Wang, M.
(2020). Invited review: Current enteric methane mitigation options. Journal of
Dairy Science, 103(7), 5759–5783. https://doi.org/10.3168/jds.2020-18906
Bennett, A. F., & Lenski, R. E. (2007). An experimental test of
evolutionary trade-offs during temperature adaptation. Proceedings of the National
Academy of Sciences, 104(Suppl 1), 8649–8654. https://doi.org/10.1073/pnas.0702117104
Berruti, A., Lumini, E., Balestrini, R., & Bianciotto, V. (2016).
Arbuscular mycorrhizal fungi as natural biofertilizers: Let’s benefit from past
successes. Frontiers in Microbiology, 6, Article 1559. https://doi.org/10.3389/fmicb.2015.01559
Bloom, A. A., Palmer, P. I., Fraser, A., Reay, D. S., & Frankenberg,
C. (2010). Large-scale controls of methanogenesis inferred from methane and gravity
spaceborne data. Science, 327(5963), 322–325. https://doi.org/10.1126/science.1175176
Bodelier, P. L. E. (2011). Interactions between nitrogenous fertilizers
and methane cycling in wetland and upland soils. Current Opinion in Environmental
Sustainability, 3(5), 379–388. https://doi.org/10.1016/j.cosust.2011.06.002
Boetius, A., & Wenzhöfer, F. (2013). Seafloor oxygen consumption
fuelled by methane from cold seeps. Nature Geoscience, 6(9), 725–734. https://doi.org/10.1038/ngeo1926
Bogner, J., Pipatti, R., Hashimoto, S., Diaz, C., Mareckova, K., Diaz,
L., Kjeldsen, P., Monni, S., Faaij, A., Gao, Q., Zhang, T., Ahmed, M. A., Sutamihardja,
R. T. M., & Gregory, R. (2008). Mitigation of global greenhouse gas emissions
from waste: Conclusions and strategies from the IPCC Fourth Assessment Report. Waste
Management & Research, 26(1), 11–32. https://doi.org/10.1177/0734242X07088433
Boyd, P. W., Claustre, H., Levy, M., Siegel, D. A., & Weber, T.
(2019). Multi-faceted particle pumps drive carbon sequestration in the ocean. Nature,
568(7752), 327–335. https://doi.org/10.1038/s41586-019-1098-2
Bradford, M. A., McCulley, R. L., Crowther, T. W., Oldfield, E. E.,
Wood, S. A., & Fierer, N. (2019). Cross-biome patterns in soil microbial respiration
predictable from evolutionary theory on thermal adaptation. Nature Ecology &
Evolution, 3(2), 223–231. https://doi.org/10.1038/s41559-018-0771-4
Bridgham, S. D., Cadillo-Quiroz, H., Keller, J. K., & Zhuang, Q.
(2013). Methane emissions from wetlands: Biogeochemical, microbial, and modeling
perspectives from local to global scales. Global Change Biology, 19(5), 1325–1346.
https://doi.org/10.1111/gcb.12131
Buesseler, K. O., Boyd, P. W., Black, E. E., & Siegel, D. A. (2020).
Metrics that matter for assessing the ocean biological carbon pump. Proceedings
of the National Academy of Sciences, 117(18), 9679–9687. https://doi.org/10.1073/pnas.1918114117
Buesseler, K. O., Lamborg, C. H., Boyd, P. W., Lam, P. J., Trull, T.
W., Bidigare, R. R., Bishop, J. K. B., Casciotti, K. L., Dehairs, F., Elskens, M.,
Honda, M., Karl, D. M., Siegel, D. A., Silver, M. W., Steinberg, D. K., Valdes,
J., Van Mooy, B., & Wilson, S. E. (2007). Revisiting carbon flux through the
ocean’s twilight zone. Science, 316(5824), 567–570. https://doi.org/10.1126/science.1137959
Carey, J. C., Tang, J., Templer, P. H., Kroeger, K. D., Crowther, T.
W., Burton, A. J., Dukes, J. S., Emmett, B., Frey, S. D., Heskel, M. A., Jiang,
L., Machmuller, M. B., Mohan, J., Panetta, A. M., Reich, P. B., Reinsch, S., Wang,
X., Allison, S. D., Bamminger, C., … Werner, J. S. (2016). Temperature response
of soil respiration largely unaltered with experimental warming. Proceedings
of the National Academy of Sciences, 113(48), 13797–13802. https://doi.org/10.1073/pnas.1605365113
Cicerone, R. J., & Oremland, R. S. (1988). Biogeochemical aspects
of atmospheric methane. Global Biogeochemical Cycles, 2(4), 299–327. https://doi.org/10.1029/GB002i004p00299
Conrad, R. (2009). The global methane cycle: Recent advances in understanding
the microbial processes involved. Environmental Microbiology Reports, 1(5),
285–292. https://doi.org/10.1111/j.1758-2229.2009.00038.x
Cotrufo, M. F., Soong, J. L., Horton, A. J., Campbell, E. E., Haddix,
M. L., Wall, D. H., & Parton, W. J. (2015). Formation of soil organic matter
via biochemical and physical pathways of litter mass loss. Nature Geoscience,
8(10), 776–779. https://doi.org/10.1038/ngeo2520
Cotrufo, M. F., Wallenstein, M. D., Boot, C. M., Denef, K., & Paul,
E. (2013). The Microbial Efficiency-Matrix Stabilization (MEMS) framework integrates
plant litter decomposition with soil organic matter stabilization: Do labile plant
inputs form stable soil organic matter? Global Change Biology, 19(4), 988–995.
https://doi.org/10.1111/gcb.12113
Crombie, A. T., & Murrell, J. C. (2014). Trace-gas metabolic versatility
of the facultative methanotroph Methylocella silvestris. Nature, 510(7503),
148–151. https://doi.org/10.1038/nature13192
Curry, C. L. (2007). Modeling the soil consumption of atmospheric methane
at the global scale. Global Biogeochemical Cycles, 21(4), GB4012. https://doi.org/10.1029/2006GB002818
Dean, J. F., Middelburg, J. J., Röckmann, T., Aerts, R., Blauw, L.
G., Egger, M., Jetten, M. S. M., de Jong, A. E. E., Meisel, O. H., Rasigraf, O.,
Slomp, C. P., in’t Zandt, M. H., & Dolman, A. J. (2018). Methane feedbacks to
the global climate system in a warmer world. Reviews of Geophysics, 56(1),
207–250. https://doi.org/10.1002/2017RG000559
Dijkstra, F. A., Prior, S. A., Runion, G. B., Torbert, H. A., Tian,
H., Lu, C., & Venterea, R. T. (2012). Effects of elevated carbon dioxide and
increased temperature on methane and nitrous oxide fluxes: Evidence from field experiments.
Frontiers in Ecology and the Environment, 10(10), 520–527. https://doi.org/10.1890/120059
Dunfield, P. F., & Knowles, R. (1995). Kinetics of inhibition of
methane oxidation by nitrate, nitrite, and ammonium in a humisol. Applied and
Environmental Microbiology, 61(8), 3129–3135. https://doi.org/10.1128/aem.61.8.3129-3135.1995
Dutaur, L., & Verchot, L. V. (2007). A global inventory of the
soil CH₄ sink. Global Biogeochemical Cycles, 21(4), GB4013. https://doi.org/10.1029/2006GB002734
Exbrayat, J.-F., Pitman, A. J., Zhang, Q., Abramowitz, G., & Wang,
Y. P. (2018). Examining soil carbon uncertainty in a global model: Response of microbial
decomposition to temperature, moisture and nutrient limitation. Biogeosciences,
15(13), 4245–4267. https://doi.org/10.5194/bg-15-4245-2018
Falkowski, P. G., Fenchel, T., & Delong, E. F. (2008). The microbial
engines that drive Earth’s biogeochemical cycles. Science, 320(5879), 1034–1039.
https://doi.org/10.1126/science.1153213
Fatichi, S., Manzoni, S., Or, D., & Paschalis, A. (2019). A mechanistic
model of microbially mediated soil biogeochemical processes: A reality check. Global
Biogeochemical Cycles, 33(6), 620–648. https://doi.org/10.1029/2018GB006077
Field, C. B., Behrenfeld, M. J., Randerson, J. T., & Falkowski,
P. (1998). Primary production of the biosphere: Integrating terrestrial and oceanic
components. Science, 281(5374), 237–240. https://doi.org/10.1126/science.281.5374.237
García-Palacios, P., McKie, B. G., Handa, I. T., Frainer, A., &
Hättenschwiler, S. (2016). The importance of litter traits and decomposers for litter
decomposition: A comparison of aquatic and terrestrial ecosystems within and across
biomes. Functional Ecology, 30(5), 819–829. https://doi.org/10.1111/1365-2435.12589
GESAMP. (2019). High level review of a wide range of proposed marine
geoengineering techniques (Rep. No. 98). IMO/FAO/UNESCO-IOC/UNIDO/WMO/IAEA/UN/UN
Environment/UNDP/ISA Joint Group of Experts on the Scientific Aspects of Marine
Environmental Protection.
Geyer, K. M., Kyker-Snowman, E., Grandy, A. S., & Frey, S. D. (2019).
Microbial carbon use efficiency: Accounting for population, community, and ecosystem-scale
controls over the fate of metabolized organic matter. Biogeochemistry, 127(2),
173–188. https://doi.org/10.1007/s10533-019-00581-6
Gómez-Garzón, C., Hernández-Santana, A., & Dussán, J. (2022). CRISPR-based
technologies for microbial community engineering. Current Opinion in Biotechnology,
73, 91–98. https://doi.org/10.1016/j.copbio.2021.07.013
Graham, E. B., Wieder, W. R., Leff, J. W., Weintraub, S. R., Townsend,
A. R., Cleveland, C. C., Philippot, L., & Nemergut, D. R. (2014). Do we need
to understand microbial communities to predict ecosystem function? A comparison
of statistical models of nitrogen cycling processes. Soil Biology and Biochemistry,
68, 279–282. https://doi.org/10.1016/j.soilbio.2013.08.023
Gupta, K., Kumar, R., Baruah, K. K., Hazarika, S., Karmakar, S., &
Bordoloi, N. (2021). Greenhouse gas emission from rice fields: A review from Indian
context. Environmental Science and Pollution Research, 28(24), 30551–30572.
https://doi.org/10.1007/s11356-021-13935-1
Hansell, D. A., Carlson, C. A., Repeta, D. J., & Schlitzer, R.
(2009). Dissolved organic matter in the ocean: A controversy stimulates new insights.
Oceanography, 22(4), 202–211. https://doi.org/10.5670/oceanog.2009.109
Henson, S. A., Sanders, R., Madsen, E., Morris, P. J., Le Moigne, F.,
& Quartly, G. D. (2011). A reduced estimate of the strength of the ocean’s biological
carbon pump. Geophysical Research Letters, 38(4), L04606. https://doi.org/10.1029/2011GL046735
Hristov, A. N., Oh, J., Lee, C., Meinen, R., Montes, F., Ott, T., Firkins,
J., Rotz, A., Dell, C., Adesogan, A., Yang, W., Tricarico, J., Kebreab, E., Waghorn,
G., Dijkstra, J., & Oosting, S. (2013). Special topics—Mitigation of methane
and nitrous oxide emissions from animal operations: I. A review of enteric methane
mitigation options. Journal of Animal Science, 91(11), 5045–5069. https://doi.org/10.2527/jas.2013-6583
Hugelius, G., Strauss, J., Zubrzycki, S., Harden, J. W., Schuur, E.
A. G., Ping, C. L., Schirrmeister, L., Grosse, G., Michaelson, G. J., Koven, C.
D., O’Donnell, J. A., Elberling, B., Mishra, U., Camill, P., Yu, Z., Palmtag, J.,
& Kuhry, P. (2014). Estimated stocks of circumpolar permafrost carbon with quantified
uncertainty ranges and identified data gaps. Biogeosciences, 11(23), 6573–6593.
https://doi.org/10.5194/bg-11-6573-2014
Intergovernmental Panel on Climate Change. (2021). Climate change
2021: The physical science basis. Contribution of Working Group I to the sixth assessment
report of the Intergovernmental Panel on Climate Change (V. Masson-Delmotte,
P. Zhai, A. Pirani, S. L. Connors, C. Péan, S. Berger, N. Caud, Y. Chen, L. Goldfarb,
M. I. Gomis, M. Huang, K. Leitzell, E. Lonnoy, J. B. R. Matthews, T. K. Maycock,
T. Waterfield, O. Yelekçi, R. Yu, & B. Zhou, Eds.). Cambridge University Press.
https://doi.org/10.1017/9781009157896
Jansson, J. K., & Hofmockel, K. S. (2020). Soil microbiomes and
climate change. Nature Reviews Microbiology, 18(1), 35–46. https://doi.org/10.1038/s41579-019-0265-7
Jiao, N., Herndl, G. J., Hansell, D. A., Benner, R., Kattner, G., Wilhelm,
S. W., Kirchman, D. L., Weinbauer, M. G., Luo, T., Chen, F., & Azam, F. (2010).
Microbial production of recalcitrant dissolved organic matter: Long-term carbon
storage in the global ocean. Nature Reviews Microbiology, 8(8), 593–599.
https://doi.org/10.1038/nrmicro2386
Jiao, N., Robinson, C., Azam, F., Thomas, H., Baltar, F., Dang, H.,
Hardman-Mountford, N. J., Johnson, M., Kirchman, D. L., Koch, B. P., Legendre, L.,
Li, C., Liu, J., Luo, T., Luo, Y. W., Mitra, A., Romanou, A., Tang, K., Wang, X.,
… Zhang, C. (2014). Mechanisms of microbial carbon sequestration in the ocean –
Future research directions. Biogeosciences, 11(19), 5285–5306. https://doi.org/10.5194/bg-11-5285-2014
Kästner, M., Miltner, A., Thiele-Bruhn, S., & Liang, C. (2021).
Microbial necromass in soils—Linking microbes to soil processes and carbon turnover.
Frontiers in Environmental Science, 9, Article 756378. https://doi.org/10.3389/fenvs.2021.756378
Knief, C. (2015). Diversity and habitat preferences of cultivated and
uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular
marker. Frontiers in Microbiology, 6, Article 1346. https://doi.org/10.3389/fmicb.2015.01346
Knittel, K., & Boetius, A. (2009). Anaerobic oxidation of methane:
Progress with an unknown process. Annual Review of Microbiology, 63, 311–334.
https://doi.org/10.1146/annurev.micro.61.080706.093130
Kolb, S., & Horn, M. A. (2012). Microbial CH₄ oxidation in acidic
soils and its importance for the global methane cycle. Environmental Microbiology
Reports, 4(3), 309–318. https://doi.org/10.1111/j.1758-2229.2012.00338.x
Kramer, M. G., & Chadwick, O. A. (2018). Climate-driven thresholds
in reactive mineral retention of soil carbon at the global scale. Nature Climate
Change, 8(12), 1104–1108. https://doi.org/10.1038/s41558-018-0341-4
Lehmann, J., Cowie, A., Masiello, C. A., Kammann, C., Woolf, D., Amonette,
J. E., Cayuela, M. L., Camps-Arbestain, M., & Whitman, T. (2021). Biochar in
climate change mitigation. Nature Geoscience, 14(12), 883–892. https://doi.org/10.1038/s41561-021-00852-8
Liang, C., Amelung, W., Lehmann, J., & Kästner, M. (2019). Quantitative
assessment of microbial necromass contribution to soil organic matter. Global
Change Biology, 25(11), 3578–3590. https://doi.org/10.1111/gcb.14781
Liang, C., Schimel, J. P., & Jastrow, J. D. (2017). The importance
of anabolism in microbial control over soil carbon storage. Nature Microbiology,
2(8), Article 17105. https://doi.org/10.1038/nmicrobiol.2017.105
Linquist, B., van Groenigen, K. J., Adviento-Borbe, M. A., Pittelkow,
C., & van Kessel, C. (2012). An agronomic assessment of greenhouse gas emissions
from major cereal crops. Global Change Biology, 18(1), 194–209. https://doi.org/10.1111/j.1365-2486.2011.02502.x
Liu, L., & Greaver, T. L. (2009). A review of nitrogen enrichment
effects on three biogenic GHGs: The CO₂ sink may be largely offset by stimulated
N₂O and CH₄ emission. Ecology Letters, 12(10), 1103–1117. https://doi.org/10.1111/j.1461-0248.2009.01351.x
Liu, S., Garcia-Palacios, P., Tedersoo, L., Guirado, E., van der Heijden,
M. G. A., Wagg, C., & Bahram, M. (2022). Plant-microbial feedbacks in soil nitrogen
cycling. Trends in Ecology & Evolution, 37(7), 599–610. https://doi.org/10.1016/j.tree.2022.03.003
MacDougall, A. H., Avis, C. A., & Weaver, A. J. (2012). Significant
contribution to climate warming from the permafrost carbon feedback. Nature Geoscience,
5(10), 719–721. https://doi.org/10.1038/ngeo1573
Machado, L., Magnusson, M., Paul, N. A., de Nys, R., & Tomkins,
N. (2018). Effects of marine and freshwater macroalgae on in vitro total gas and
methane production. PLOS ONE, 13(5), e0197423. https://doi.org/10.1371/journal.pone.0197423
Manzoni, S., Taylor, P., Richter, A., Porporato, A., & Ågren, G.
I. (2012). Environmental and stoichiometric controls on microbial carbon-use efficiency
in soils. New Phytologist, 196(1), 79–91. https://doi.org/10.1111/j.1469-8137.2012.04225.x
Martin, J. H., Knauer, G. A., Karl, D. M., & Broenkow, W. W. (1987).
VERTEX: Carbon cycling in the northeast Pacific. Deep Sea Research Part A. Oceanographic
Research Papers, 34(2), 267–285. https://doi.org/10.1016/0198-0149(87)90086-0
McGlynn, S. E. (2017). Energy metabolism during anaerobic methane oxidation
in ANME archaea. Microbes and Environments, 32(1), 5–13. https://doi.org/10.1264/jsme2.ME16166
Mohan, J. E., Cowden, C. C., Baas, P., Dawadi, A., Frankson, P. T.,
Helmick, K., & Witt, C. A. (2014). Mycorrhizal fungi mediation of terrestrial
ecosystem responses to global change: Mini-review. Fungal Ecology, 10, 3–19.
https://doi.org/10.1016/j.funeco.2014.01.005
Moomaw, W. R., Chmura, G. L., Davies, G. T., Finlayson, C. M., Middleton,
B. A., Natali, S. M., Perry, J. E., Roulet, N., & Sutton-Grier, A. E. (2018).
Wetlands in a changing climate: Science, policy and management. Wetlands, 38(2),
183–205. https://doi.org/10.1007/s13157-018-1023-8
Mopper, K., Stubbins, A., Ritchie, J. D., Bialk, H. M., & Hatcher,
P. G. (2015). Advanced instrumental approaches for characterization of marine dissolved
organic matter: Extraction techniques, mass spectrometry, and nuclear magnetic resonance
spectroscopy. Chemical Reviews, 107(2), 419–442. https://doi.org/10.1021/cr050359b
Nisbet, E. G., Manning, M. R., Dlugokencky, E. J., Fisher, R. E., Lowry,
D., Michel, S. E., Myhre, C. L., Platt, S. M., Allen, G., Bousquet, P., Brownlow,
R., Cain, M., France, J. L., Hermansen, O., Hossaini, R., Jones, A. E., Levin, I.,
Manning, A. C., Myhre, G., … White, J. W. C. (2019). Very strong atmospheric methane
growth in the 4 years 2014–2017: Implications for the Paris Agreement. Global
Biogeochemical Cycles, 33(3), 318–342. https://doi.org/10.1029/2018GB006009
Paustian, K., Lehmann, J., Ogle, S., Reay, D., Robertson, G. P., &
Smith, P. (2016). Climate-smart soils. Nature, 532(7597), 49–57. https://doi.org/10.1038/nature17174
Raich, J. W., & Schlesinger, W. H. (1992). The global carbon dioxide
flux in soil respiration and its relationship to vegetation and climate. Tellus
B: Chemical and Physical Meteorology, 44(2), 81–99. https://doi.org/10.1034/j.1600-0889.1992.t01-1-00001.x
Regnier, P., Dale, A. W., Arndt, S., LaRowe, D. E., Mogollón, J., &
Van Cappellen, P. (2011). Quantitative analysis of anaerobic oxidation of methane
(AOM) in marine sediments: A modeling perspective. *Earth-Science Reviews, 106*(1–2),
105–130. https://doi.org/10.1016/j.earscirev.2011.01.002
Riley, W. J., Subin, Z. M., Lawrence, D. M., Swenson, S. C., Torn,
M. S., Meng, L., Mahowald, N. M., & Hess, P. (2011). Barriers to predicting
changes in global terrestrial methane fluxes: Analyses using CLM4Me, a methane biogeochemistry
model integrated in CESM. Biogeosciences, 8(7), 1925–1953. https://doi.org/10.5194/bg-8-1925-2011
Robinson, C., Bennett, C., Bliss, J., García-Martín, E., Gardner, J.,
& Ng, M. (2015). Interactions between the marine biogeochemical cycles of
carbon, nitrogen and phosphorus [Figure]. Figshare. https://doi.org/10.6084/m9.figshare.1585741.v1
Roque, B. M., Venegas, M., Kinley, R. D., de Nys, R., Duarte, T. L.,
Yang, X., & Kebreab, E. (2021). Red seaweed (Asparagopsis taxiformis) supplementation
reduces enteric methane by over 80 percent in beef steers. PLOS ONE, 16(3),
e0247820. https://doi.org/10.1371/journal.pone.0247820
Rosentreter, J. A., Borges, A. V., Deemer, B. R., Holgerson, M. A.,
Liu, S., Song, C., Melack, J., Raymond, P. A., Duarte, C. M., Allen, G. H., Olefeldt,
D., Poulter, B., Battin, T. I., & Eyre, B. D. (2021). Half of global methane
emissions come from highly variable aquatic ecosystem sources. Nature Geoscience,
14(4), 225–230. https://doi.org/10.1038/s41561-021-00715-2
Ruser, R., & Schulz, R. (2015). The effect of nitrification inhibitors
on the nitrous oxide (N₂O) release from agricultural soils—A review. Journal
of Plant Nutrition and Soil Science, 178(2), 171–188. https://doi.org/10.1002/jpln.201400251
Sander, B. O., Samson, M., & Buresh, R. J. (2014). Methane and
nitrous oxide emissions from flooded rice fields as affected by water and straw
management between rice crops. Geoderma, 235–236, 355–362. https://doi.org/10.1016/j.geoderma.2014.07.020
Sanderson, M. G. (1996). Biomass of termites and their emissions of
methane and carbon dioxide: A global database. Global Biogeochemical Cycles,
10(4), 543–557. https://doi.org/10.1029/96GB01933
Saunois, M., Stavert, A. R., Poulter, B., Bousquet, P., Canadell, J.
G., Jackson, R. B., Raymond, P. A., Dlugokencky, E. J., Houweling, S., Patra, P.
K., Ciais, P., Arora, V. K., Bastviken, D., Bergamaschi, P., Blake, D. R., Brailsford,
G., Bruhwiler, L., Carlson, K. M., Carrol, M., … Zhuang, Q. (2020). The Global Methane
Budget 2000–2017. Earth System Science Data, 12(3), 1561–1623. https://doi.org/10.5194/essd-12-1561-2020
Schädel, C., Bader, M. K.-F., Schuur, E. A. G., Biasi, C., Bracho,
R., Čapek, P., De Baets, S., Diáková, K., Ernakovich, J., Estop-Aragonés, C., Graham,
D. E., Hartley, I. P., Iversen, C. M., Kane, E., Knoblauch, C., Lupascu, M., Martikainen,
P. J., Natali, S. M., Norby, R. J., … Wickland, K. P. (2016). Potential carbon emissions
dominated by carbon dioxide from thawed permafrost soils. Nature Climate Change,
6(10), 950–953. https://doi.org/10.1038/nclimate3054
Schaefer, K., Lantuit, H., Romanovsky, V. E., Schuur, E. A. G., &
Witt, R. (2014). The impact of the permafrost carbon feedback on global climate.
Environmental Research Letters, 9(8), Article 085003. https://doi.org/10.1088/1748-9326/9/8/085003
Scheffer, M., Bascompte, J., Brock, W. A., Brovkin, V., Carpenter,
S. R., Dakos, V., Held, H., van Nes, E. H., Rietkerk, M., & Sugihara, G. (2009).
Early-warning signals for critical transitions. Nature, 461(7260), 53–59.
https://doi.org/10.1038/nature08227
Scheutz, C., Kjeldsen, P., Bogner, J. E., De Visscher, A., Gebert,
J., Hilger, H. A., Huber-Humer, M., & Spokas, K. (2009). Microbial methane oxidation
processes and technologies for mitigation of landfill gas emissions. Waste Management
& Research, 27(5), 409–455. https://doi.org/10.1177/0734242X09339325
Schmidtko, S., Stramma, L., & Visbeck, M. (2017). Decline in global
oceanic oxygen content during the past five decades. Nature, 542(7641), 335–339.
https://doi.org/10.1038/nature21399
Schuur, E. A. G., McGuire, A. D., Schädel, C., Grosse, G., Harden,
J. W., Hayes, D. J., Hugelius, G., Koven, C. D., Kuhry, P., Lawrence, D. M., Natali,
S. M., Olefeldt, D., Romanovsky, V. E., Schaefer, K., Turetsky, M. R., Treat, C.
C., & Vonk, J. E. (2015). Climate change and the permafrost carbon feedback.
Nature, 520(7546), 171–179. https://doi.org/10.1038/nature14338
Schwander, T., Schada von Borzyskowski, L., Burgener, S., Cortina,
N. S., & Erb, T. J. (2016). A synthetic pathway for the fixation of carbon dioxide
in vitro. Science, 354(6314), 900–904. https://doi.org/10.1126/science.aah5237
Schwietzke, S., Sherwood, O. A., Bruhwiler, L. M. P., Miller, J. B.,
Etiope, G., Dlugokencky, E. J., Michel, S. E., Arling, V. A., Vaughn, B. H., White,
J. W. C., & Tans, P. P. (2016). Upward revision of global fossil fuel methane
emissions based on isotope database. Nature, 538(7623), 88–91. https://doi.org/10.1038/nature19797
Sinsabaugh, R. L., Manzoni, S., Moorhead, D. L., & Richter, A.
(2013). Carbon use efficiency of microbial communities: Stoichiometry, methodology
and modelling. Ecology Letters, 16(7), 775–784. https://doi.org/10.1111/ele.12113
Smith, K. A., Dobbie, K. E., Ball, B. C., Bakken, L. R., Sitaula, B.
K., Hansen, S., Brumme, R., Borken, W., Christensen, S., Priemé, A., Fowler, D.,
MacDonald, J. A., Skiba, U., Klemedtsson, L., Kasimir-Klemedtsson, A., Degórska,
A., & Orlanski, P. (2000). Oxidation of atmospheric methane in Northern European
soils, comparison with other ecosystems, and uncertainties in the global terrestrial
sink. Global Change Biology, 6(7), 791–803. https://doi.org/10.1046/j.1365-2486.2000.00356.x
Suttle, C. A. (2007). Marine viruses—Major players in the global ecosystem.
Nature Reviews Microbiology, 5(10), 801–812. https://doi.org/10.1038/nrmicro1750
Turetsky, M. R., Abbott, B. W., Jones, M. C., Anthony, K. W., Olefeldt,
D., Schuur, E. A. G., Grosse, G., Kuhry, P., Hugelius, G., Koven, C., Lawrence,
D. M., Gibson, C., Sannel, A. B. K., & McGuire, A. D. (2020). Carbon release
through abrupt permafrost thaw. Nature Geoscience, 13(2), 138–143. https://doi.org/10.1038/s41561-019-0526-0
Turetsky, M. R., Benscoter, B., Page, S., Rein, G., van der Werf, G.
R., & Watts, A. (2015). Global vulnerability of peatlands to fire and carbon
loss. Nature Geoscience, 8(1), 11–14. https://doi.org/10.1038/ngeo2325
Turetsky, M. R., Kotowska, A., Bubier, J., Dise, N. B., Crill, P.,
Hornibrook, E. R. C., Minkkinen, K., Moore, T. R., Myers-Smith, I. H., Nykänen,
H., Olefeldt, D., Rinne, J., Saarnio, S., Shurpali, N., Tuittila, E. S., Waddington,
J. M., White, J. R., Wickland, K. P., & Wilmking, M. (2014). A synthesis of
methane emissions from 71 northern, temperate, and subtropical wetlands. Global
Change Biology, 20(7), 2183–2197. https://doi.org/10.1111/gcb.12580
van Gestel, N., Shi, Z., van Groenigen, K. J., Osenberg, C. W., Andresen,
L. C., Dukes, J. S., Hovenden, M. J., Luo, Y., Michelsen, A., Pendall, E., Reich,
P. B., Schuur, E. A. G., & Hungate, B. A. (2018). Predicting soil carbon loss
with warming. Nature, 554(7693), E4–E5. https://doi.org/10.1038/nature25745
Voigt, C., Marushchak, M. E., Lamprecht, R. E., Jackowicz-Korczyński,
M., Lindgren, A., Masterpanov, M., Granlund, L., Christensen, T. R., Tahvanainen,
T., Martikainen, P. J., & Biasi, C. (2017). Increased nitrous oxide emissions
from Arctic peatlands after permafrost thaw. Proceedings of the National Academy
of Sciences, 114(24), 6238–6243. https://doi.org/10.1073/pnas.1702902114
Walter Anthony, K., Daanen, R., Anthony, P., Schneider von Deimling,
T., Ping, C.-L., Chanton, J. P., & Grosse, G. (2016). Methane emissions proportional
to permafrost carbon thawed in Arctic lakes since the 1950s. Nature Geoscience,
9(9), 679–682. https://doi.org/10.1038/ngeo2795
Wang, F., Harindintwali, J.-D., Wei, K., Wang, Z., Li, B., & et
al. (2023). Climate change: Strategies for mitigation and adaptation. The Innovation
Geoscience, 1(1), 100015. https://doi.org/10.59717/j.xinn-geo.2023.100015
Wang, G., Jagadamma, S., Mayes, M. A., Schadt, C. W., Steinweg, J.
M., Gu, L., & Post, W. M. (2021). Microbial dormancy improves development and
experimental validation of ecosystem model. The ISME Journal, 15(4), 1145–1162.
https://doi.org/10.1038/s41396-020-00845-2
Ward, N., Larsen, Ø., Sakwa, J., Bruseth, L., Khouri, H., Scott Durkin,
A., & et al. (2015). Global Methane Cycle. PLOS Biology. [Figure]. https://doi.org/10.1371/journal.pbio.0020303.g001
Weber, T., Wiseman, N. A., & Kock, A. (2019). Global ocean methane
emissions dominated by shallow coastal waters. Nature Communications, 10(1),
Article 4584. https://doi.org/10.1038/s41467-019-12541-7
Weinbauer, M. G., Bettarel, Y., Cattaneo, R., Luef, B., Maier, C.,
Motegi, C., Peduzzi, P., & Mari, X. (2011). Viral ecology of organic and inorganic
particles in aquatic systems: Avenues for further research. Aquatic Microbial
Ecology, 64(1), 1–20. https://doi.org/10.3354/ame01506
Wieder, W. R., Allison, S. D., Davidson, E. A., Georgiou, K., Hararuk,
O., He, Y., Hopkins, F., Luo, Y., Smith, M. J., Sulman, B., Todd-Brown, K., Wang,
Y. P., Xia, J., & Xu, X. (2015). Explicitly representing soil microbial processes
in Earth system models. Global Biogeochemical Cycles, 29(10), 1782–1800.
https://doi.org/10.1002/2015GB005188
Wieder, W. R., Bonan, G. B., & Allison, S. D. (2013). Global soil
carbon projections are improved by modelling microbial processes. Nature Climate
Change, 3(10), 909–912. https://doi.org/10.1038/nclimate1951
Wilson, R. M., Hopple, A. M., Tfaily, M. M., Sebestyen, S. D., Schadt,
C. W., Pfeifer-Meister, L., Medvedeff, C., McFarlane, K. J., Kostka, J. E., Kolton,
M., Chanton, J. P., Cooper, W. T., Bridgham, S. D., & Hanson, P. J. (2016).
Stability of peatland carbon to rising temperatures. Nature Communications, 7,
Article 13723. https://doi.org/10.1038/ncomms13723
Yvon-Durocher, G., Allen, A. P., Bastviken, D., Conrad, R., Gudasz,
C., St-Pierre, A., Thanh-Duc, N., & del Giorgio, P. A. (2014). Methane fluxes
show consistent temperature dependence across microbial to ecosystem scales. Nature,
507(7493), 488–491. https://doi.org/10.1038/nature13164
Zhang, Z., Zimmermann, N. E., Stenke, A., Li, X., Hodson, E. L., Zhu,
G., Huang, C., & Poulter, B. (2017). Emerging role of wetland methane emissions
in driving 21st century climate change. Proceedings of the National Academy of
Sciences, 114(36), 9647–9652. https://doi.org/10.1073/pnas.1618765114
Zimmerman, A. E., Howard-Varona, C., Needham, D. M., John, S. G., Worden,
A. Z., Sullivan, M. B., Weitz, J. S., & Waldbauer, J. R. (2020). Metabolic and
biogeochemical consequences of viral infection in aquatic ecosystems. Nature
Reviews Microbiology, 18(1), 21–34. https://doi.org/10.1038/s41579-019-0270-x
Wilcox, M. H., et al. (2017). Bezlotoxumab
for prevention of recurrent Clostridium difficile infection. New England Journal
of Medicine, 376 (4), 305-317. https://doi.org/10.1056/NEJMoa1602615